SARS CoV-2 Mpro 1-302 C145A in complex with peptide 7
Liu, M., Huang, H.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Replicase polyprotein 1ab | 302 | Severe acute respiratory syndrome coronavirus 2 | Mutation(s): 0 Gene Names: rep, 1a-1b EC: 3.4.22.69 | 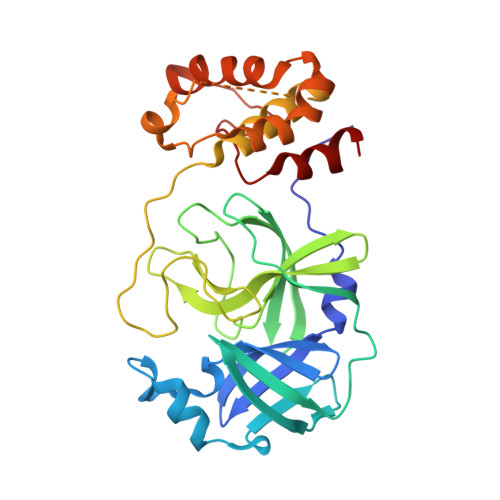 | |
UniProt | |||||
Find proteins for P0DTD1 (Severe acute respiratory syndrome coronavirus 2) Explore P0DTD1 Go to UniProtKB: P0DTD1 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0DTD1 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Find similar proteins by: Sequence | 3D Structure
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| VAL-LYS-LEU-GLN-ALA-VAL-PHE-ARG | 8 | Severe acute respiratory syndrome coronavirus 2 | Mutation(s): 0 | 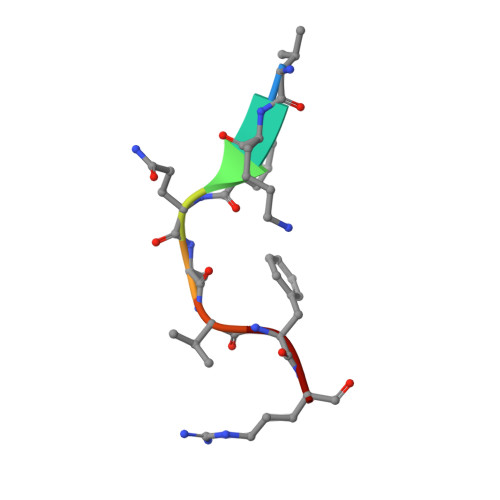 | |
Sequence AnnotationsExpand | |||||
| |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 66.892 | α = 90 |
| b = 66.892 | β = 90 |
| c = 240.659 | γ = 120 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| SCALA | data scaling |
| PDB_EXTRACT | data extraction |
| CrysalisPro | data reduction |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Not funded | -- |