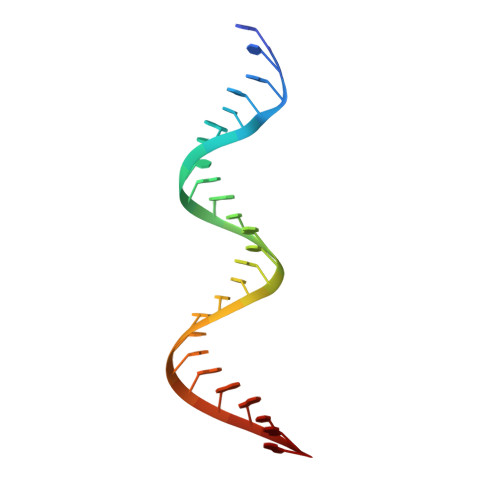
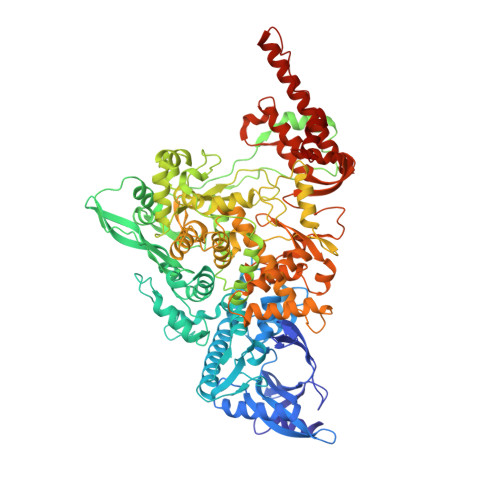
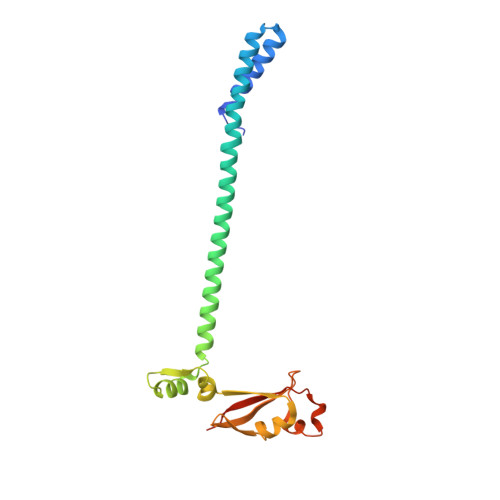
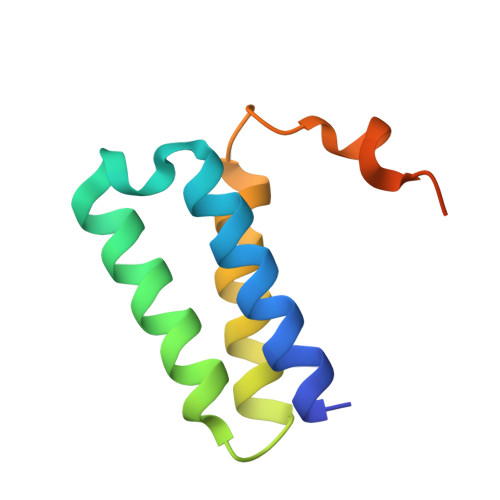
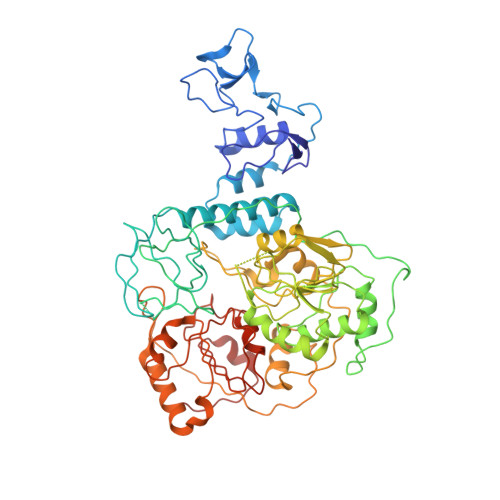
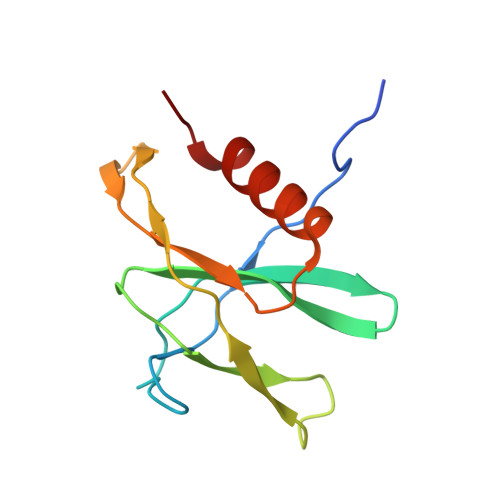
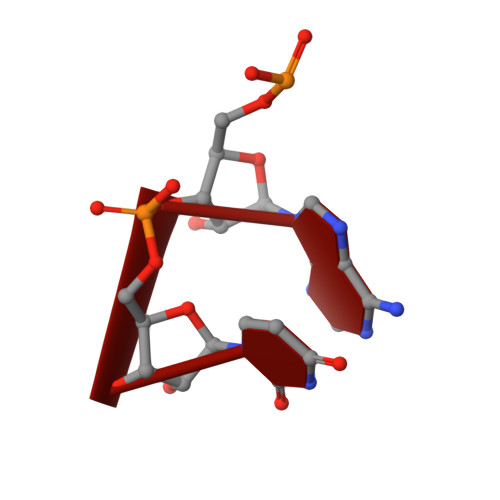
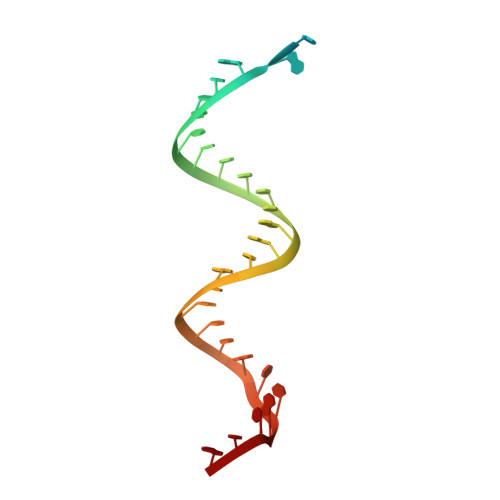
A mechanism for SARS-CoV-2 RNA capping and its inhibition by nucleotide analog inhibitors.
Yan, L., Huang, Y., Ge, J., Liu, Z., Lu, P., Huang, B., Gao, S., Wang, J., Tan, L., Ye, S., Yu, F., Lan, W., Xu, S., Zhou, F., Shi, L., Guddat, L.W., Gao, Y., Rao, Z., Lou, Z.(2022) Cell 185: 4347-4360.e17
- PubMed: 36335936
- DOI: https://doi.org/10.1016/j.cell.2022.09.037
- Primary Citation of Related Structures:
8GW1, 8GWB, 8GWE, 8GWF, 8GWG, 8GWI, 8GWK, 8GWM, 8GWN, 8GWO - PubMed Abstract:
Decoration of cap on viral RNA plays essential roles in SARS-CoV-2 proliferation. Here, we report a mechanism for SARS-CoV-2 RNA capping and document structural details at atomic resolution. The NiRAN domain in polymerase catalyzes the covalent link of RNA 5' end to the first residue of nsp9 (termed as RNAylation), thus being an intermediate to form cap core (GpppA) with GTP catalyzed again by NiRAN. We also reveal that triphosphorylated nucleotide analog inhibitors can be bonded to nsp9 and fit into a previously unknown "Nuc-pocket" in NiRAN, thus inhibiting nsp9 RNAylation and formation of GpppA. S-loop (residues 50-KTN-52) in NiRAN presents a remarkable conformational shift observed in RTC bound with sofosbuvir monophosphate, reasoning an "induce-and-lock" mechanism to design inhibitors. These findings not only improve the understanding of SARS-CoV-2 RNA capping and the mode of action of NAIs but also provide a strategy to design antiviral drugs.
Organizational Affiliation:
MOE Key Laboratory of Protein Science, School of Medicine, Tsinghua University, Beijing, China.