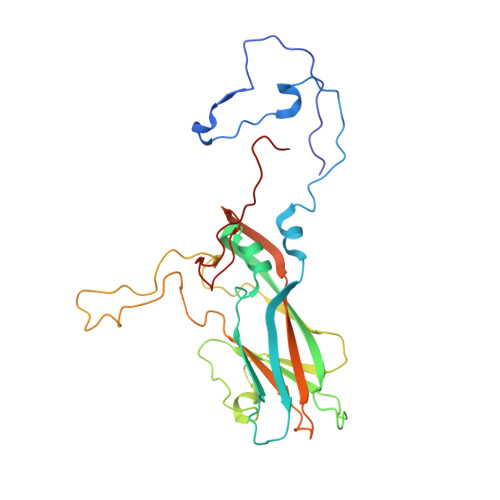
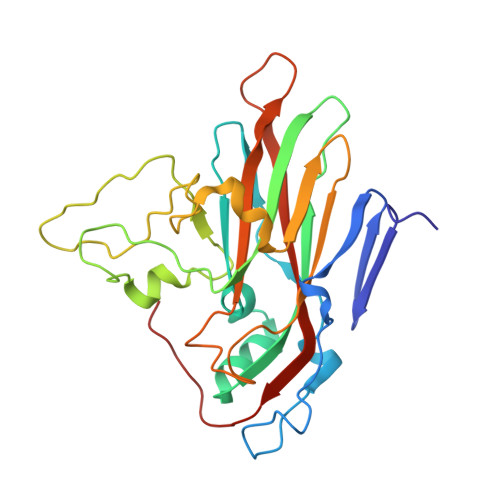
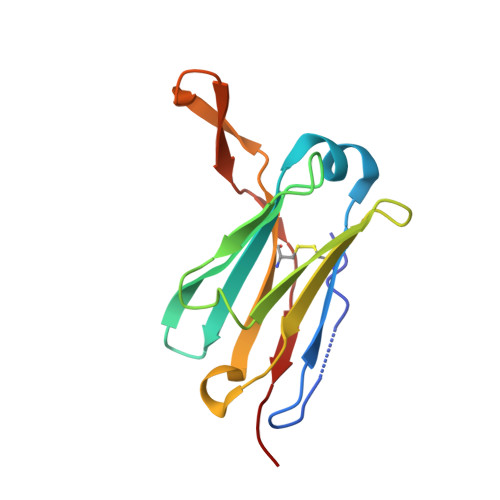
A human monoclonal antibody binds within the poliovirus receptor-binding site to neutralize all three serotypes.
Charnesky, A.J., Faust, J.E., Lee, H., Puligedda, R.D., Goetschius, D.J., DiNunno, N.M., Thapa, V., Bator, C.M., Cho, S.H.J., Wahid, R., Mahmood, K., Dessain, S., Chumakov, K.M., Rosenfeld, A., Hafenstein, S.L.(2023) Nat Commun 14: 6335-6335
- PubMed: 37816742
- DOI: https://doi.org/10.1038/s41467-023-41052-9
- Primary Citation of Related Structures:
8E8L, 8E8R, 8E8S, 8E8X, 8E8Y, 8E8Z - PubMed Abstract:
Global eradication of poliovirus remains elusive, and it is critical to develop next generation vaccines and antivirals. In support of this goal, we map the epitope of human monoclonal antibody 9H2 which is able to neutralize the three serotypes of poliovirus. Using cryo-EM we solve the near-atomic structures of 9H2 fragments (Fab) bound to capsids of poliovirus serotypes 1, 2, and 3. The Fab-virus complexes show that Fab interacts with the same binding mode for each serotype and at the same angle of interaction relative to the capsid surface. For each of the Fab-virus complexes, we find that the binding site overlaps with the poliovirus receptor (PVR) binding site and maps across and into a depression in the capsid called the canyon. No conformational changes to the capsid are induced by Fab binding for any complex. Competition binding experiments between 9H2 and PVR reveal that 9H2 impedes receptor binding. Thus, 9H2 outcompetes the receptor to neutralize poliovirus. The ability to neutralize all three serotypes, coupled with the critical importance of the conserved receptor binding site make 9H2 an attractive antiviral candidate for future development.
Organizational Affiliation:
Molecular, Cellular, and Integrative Biosciences Program, The Pennsylvania State University, University Park, PA, USA.