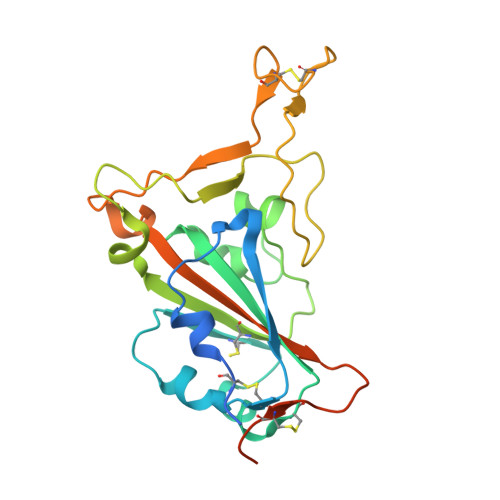
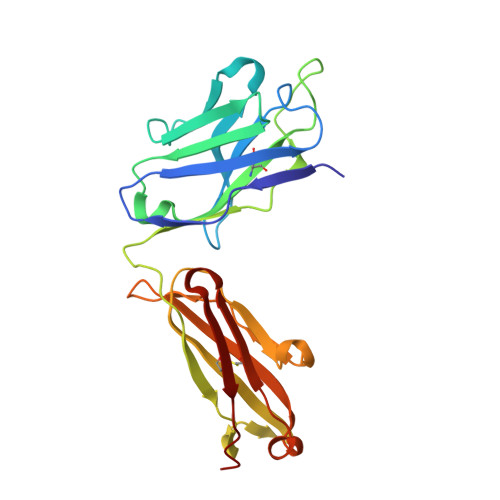
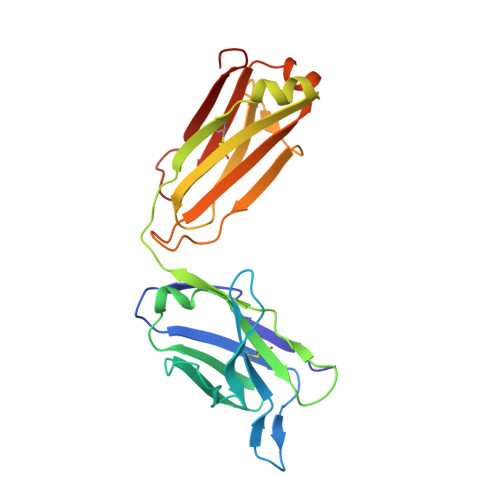
A multi-specific, multi-affinity antibody platform neutralizes sarbecoviruses and confers protection against SARS-CoV-2 in vivo.
Burn Aschner, C., Muthuraman, K., Kucharska, I., Cui, H., Prieto, K., Nair, M.S., Wang, M., Huang, Y., Christie-Holmes, N., Poon, B., Lam, J., Sultana, A., Kozak, R., Mubareka, S., Rubinstein, J.L., Rujas, E., Treanor, B., Ho, D.D., Jetha, A., Julien, J.P.(2023) Sci Transl Med 15: eadf4549-eadf4549
- PubMed: 37224226
- DOI: https://doi.org/10.1126/scitranslmed.adf4549
- Primary Citation of Related Structures:
8DNN - PubMed Abstract:
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the causative agent of coronavirus disease 2019 (COVID-19), has been responsible for a global pandemic. Monoclonal antibodies (mAbs) have been used as antiviral therapeutics; however, these therapeutics have been limited in efficacy by viral sequence variability in emerging variants of concern (VOCs) and in deployment by the need for high doses. In this study, we leveraged the multi-specific, multi-affinity antibody (Multabody, MB) platform, derived from the human apoferritin protomer, to enable the multimerization of antibody fragments. MBs were shown to be highly potent, neutralizing SARS-CoV-2 at lower concentrations than their corresponding mAb counterparts. In mice infected with SARS-CoV-2, a tri-specific MB targeting three regions within the SARS-CoV-2 receptor binding domain was protective at a 30-fold lower dose than a cocktail of the corresponding mAbs. Furthermore, we showed in vitro that mono-specific MBs potently neutralize SARS-CoV-2 VOCs by leveraging augmented avidity, even when corresponding mAbs lose their ability to neutralize potently, and that tri-specific MBs expanded the neutralization breadth beyond SARS-CoV-2 to other sarbecoviruses. Our work demonstrates how avidity and multi-specificity combined can be leveraged to confer protection and resilience against viral diversity that exceeds that of traditional monoclonal antibody therapies.
Organizational Affiliation:
Program in Molecular Medicine, Hospital for Sick Children Research Institute, Toronto, ON M5G 0A4, Canada.