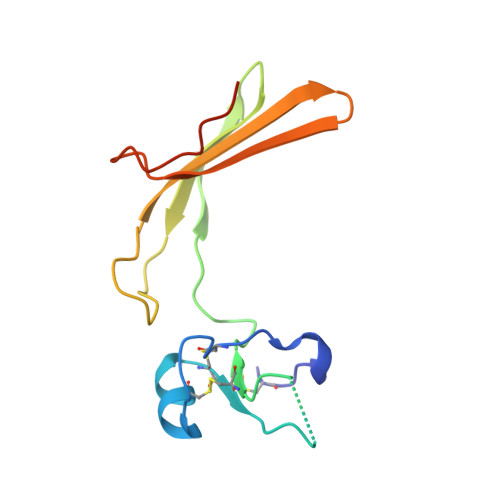
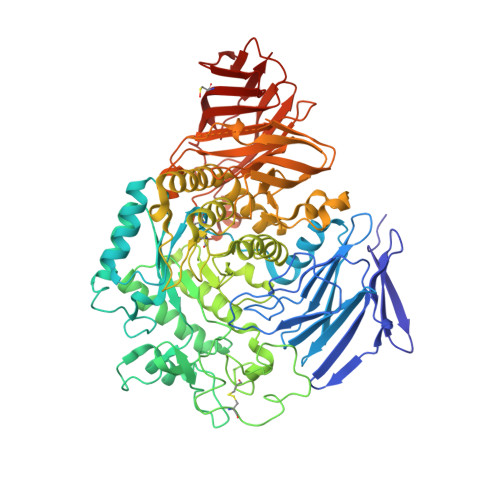
Fluorescence polarisation activity-based protein profiling for the identification of deoxynojirimycin-type inhibitors selective for lysosomal retaining alpha- and beta-glucosidases.
van der Gracht, D., Rowland, R.J., Roig-Zamboni, V., Ferraz, M.J., Louwerse, M., Geurink, P.P., Aerts, J.M.F.G., Sulzenbacher, G., Davies, G.J., Overkleeft, H.S., Artola, M.(2023) Chem Sci 14: 9136-9144
- PubMed: 37655021
- DOI: https://doi.org/10.1039/d3sc01021j
- Primary Citation of Related Structures:
7NWV, 8CB1, 8CB6 - PubMed Abstract:
Lysosomal exoglycosidases are responsible for processing endocytosed glycans from the non-reducing end to produce the corresponding monosaccharides. Genetic mutations in a particular lysosomal glycosidase may result in accumulation of its particular substrate, which may cause diverse lysosomal storage disorders. The identification of effective therapeutic modalities to treat these diseases is a major yet poorly realised objective in biomedicine. One common strategy comprises the identification of effective and selective competitive inhibitors that may serve to stabilize the proper folding of the mutated enzyme, either during maturation and trafficking to, or residence in, endo-lysosomal compartments. The discovery of such inhibitors is greatly aided by effective screening assays, the development of which is the focus of the here-presented work. We developed and applied fluorescent activity-based probes reporting on either human GH30 lysosomal glucosylceramidase (GBA1, a retaining β-glucosidase) or GH31 lysosomal retaining α-glucosidase (GAA). FluoPol-ABPP screening of our in-house 358-member iminosugar library yielded compound classes selective for either of these enzymes. In particular, we identified a class of N -alkyldeoxynojirimycins that inhibit GAA, but not GBA1, and that may form the starting point for the development of pharmacological chaperone therapeutics for the lysosomal glycogen storage disease that results from genetic deficiency in GAA: Pompe disease.
Organizational Affiliation:
Leiden Institute of Chemistry, Leiden University P. O. Box 9502 2300 RA Leiden The Netherlands h.s.overkleeft@lic.leidenuniv.nl m.e.artola@lic.leidenuniv.nl.