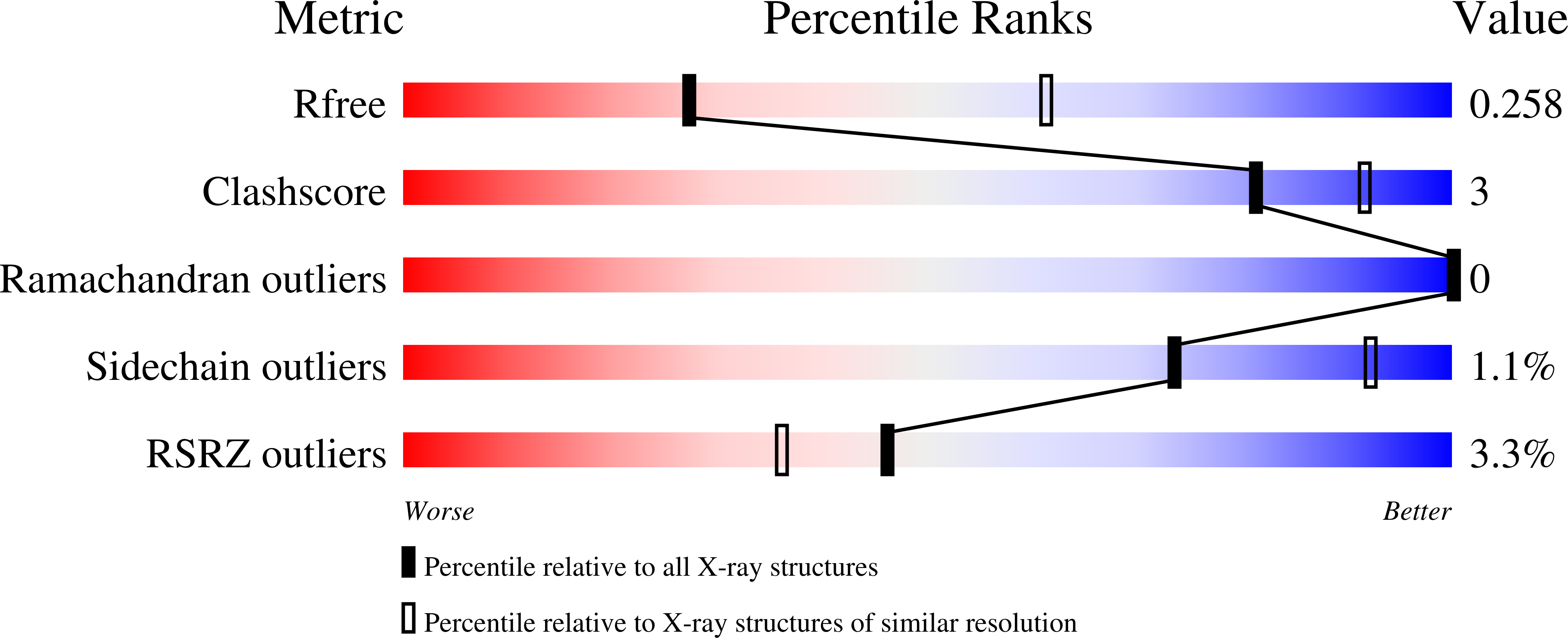
Structural basis for selective modification of Rho and Ras GTPases by Clostridioides difficile toxin B.
Liu, Z., Zhang, S., Chen, P., Tian, S., Zeng, J., Perry, K., Dong, M., Jin, R.(2021) Sci Adv 7: eabi4582-eabi4582
- PubMed: 34678063
- DOI: https://doi.org/10.1126/sciadv.abi4582
- Primary Citation of Related Structures:
7S0Y, 7S0Z - PubMed Abstract:
Toxin B (TcdB) is a primary cause of Clostridioides difficile infection (CDI). This toxin acts by glucosylating small GTPases in the Rho/Ras families, but the structural basis for TcdB recognition and selectivity of specific GTPase substrates remain unsolved. Here, we report the cocrystal structures of the glucosyltransferase domain (GTD) of two distinct TcdB variants in complex with human Cdc42 and R-Ras, respectively. These structures reveal a common structural mechanism by which TcdB recognizes Rho and R-Ras. Furthermore, we find selective clustering of adaptive residue changes in GTDs that determine their substrate preferences, which helps partition all known TcdB variants into two groups that display distinct specificities toward Rho or R-Ras. Mutations that selectively disrupt GTPases binding reduce the glucosyltransferase activity of the GTD and the toxicity of TcdB holotoxin. These findings establish the structural basis for TcdB recognition of small GTPases and reveal strategies for therapeutic interventions for CDI.
Organizational Affiliation:
Department of Physiology and Biophysics, University of California, Irvine, Irvine, CA 92697, USA.