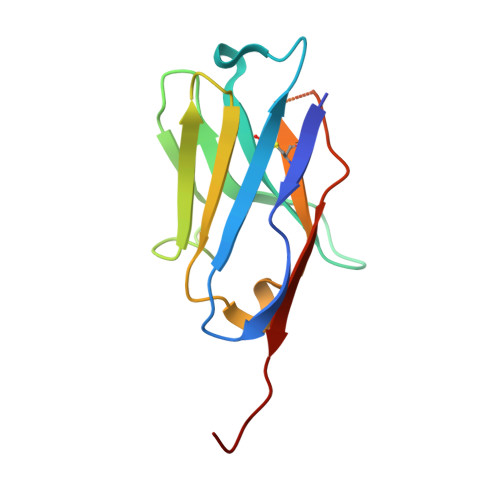
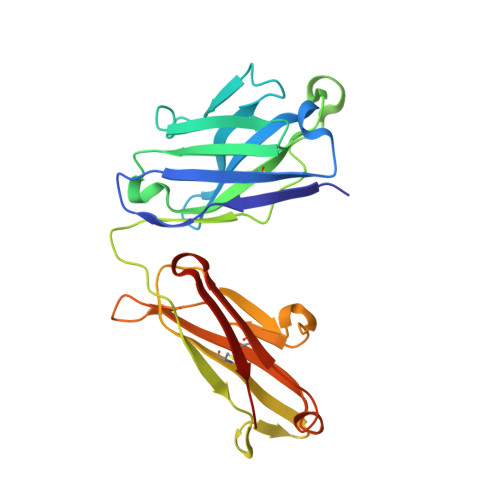
Cryo-EM structure determination of small proteins by nanobody-binding scaffolds (Legobodies).
Wu, X., Rapoport, T.A.(2021) Proc Natl Acad Sci U S A 118
- PubMed: 34620716
- DOI: https://doi.org/10.1073/pnas.2115001118
- Primary Citation of Related Structures:
7R9D, 7RXC, 7RXD - PubMed Abstract:
We describe a general method that allows structure determination of small proteins by single-particle cryo-electron microscopy (cryo-EM). The method is based on the availability of a target-binding nanobody, which is then rigidly attached to two scaffolds: 1) a Fab fragment of an antibody directed against the nanobody and 2) a nanobody-binding protein A fragment fused to maltose binding protein and Fab-binding domains. The overall ensemble of ∼120 kDa, called Legobody, does not perturb the nanobody-target interaction, is easily recognizable in EM images due to its unique shape, and facilitates particle alignment in cryo-EM image processing. The utility of the method is demonstrated for the KDEL receptor, a 23-kDa membrane protein, resulting in a map at 3.2-Å overall resolution with density sufficient for de novo model building, and for the 22-kDa receptor-binding domain (RBD) of SARS-CoV-2 spike protein, resulting in a map at 3.6-Å resolution that allows analysis of the binding interface to the nanobody. The Legobody approach thus overcomes the current size limitations of cryo-EM analysis.
Organizational Affiliation:
HHMI, Harvard Medical School, Boston, MA 02115; tom_rapoport@hms.harvard.edu xudong_wu2@hms.harvard.edu.