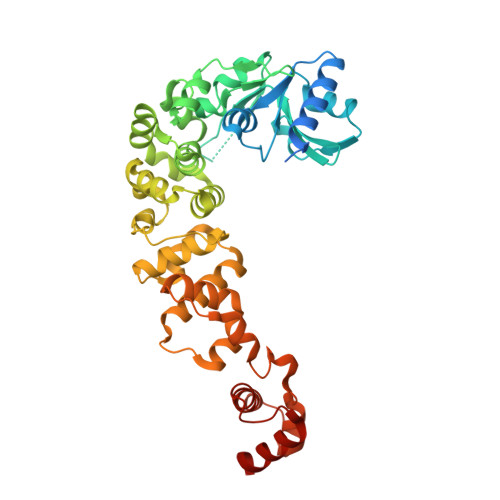
CCA-addition in the cold: Structural characterization of the psychrophilic CCA-adding enzyme from the permafrost bacterium Planococcus halocryophilus .
de Wijn, R., Rollet, K., Ernst, F.G.M., Wellner, K., Betat, H., Morl, M., Sauter, C.(2021) Comput Struct Biotechnol J 19: 5845-5855
- PubMed: 34765099
- DOI: https://doi.org/10.1016/j.csbj.2021.10.018
- Primary Citation of Related Structures:
7OQX, 7OTL, 7OTR - PubMed Abstract:
CCA-adding enzymes are highly specific RNA polymerases that add and maintain the sequence C-C-A at tRNA 3'-ends. Recently, we could reveal that cold adaptation of such a polymerase is not only achieved at the expense of enzyme stability, but also at the cost of polymerization fidelity. Enzymes from psychrophilic organisms usually show an increased structural flexibility to enable catalysis at low temperatures. Here, polymerases face a dilemma, as there is a discrepancy between the need for a tightly controlled flexibility during polymerization and an increased flexibility as strategy for cold adaptation. Based on structural and biochemical analyses, we contribute to clarify the cold adaptation strategy of the psychrophilic CCA-adding enzyme from Planococcus halocryophilus , a gram-positive bacterium thriving in the arctic permafrost at low temperatures down to -15 °C. A comparison with the closely related enzyme from the thermophilic bacterium Geobacillus stearothermophilus reveals several features of cold adaptation - a significantly reduced amount of alpha-helical elements in the C-terminal tRNA-binding region and a structural adaptation in one of the highly conserved catalytic core motifs located in the N-terminal catalytic core of the enzyme.
Organizational Affiliation:
Architecture et Réactivité de l'ARN, Université de Strasbourg, CNRS, IBMC, 67084 Strasbourg, France.