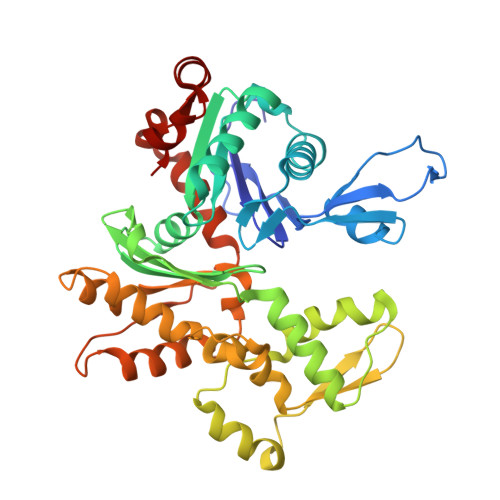
Higher-order phosphatase-substrate contacts terminate the integrated stress response.
Yan, Y., Harding, H.P., Ron, D.(2021) Nat Struct Mol Biol 28: 835-846
- PubMed: 34625748
- DOI: https://doi.org/10.1038/s41594-021-00666-7
- Primary Citation of Related Structures:
7NXV, 7NZM - PubMed Abstract:
Many regulatory PPP1R subunits join few catalytic PP1c subunits to mediate phosphoserine and phosphothreonine dephosphorylation in metazoans. Regulatory subunits engage the surface of PP1c, locally affecting flexible access of the phosphopeptide to the active site. However, catalytic efficiency of holophosphatases towards their phosphoprotein substrates remains unexplained. Here we present a cryo-EM structure of the tripartite PP1c-PPP1R15A-G-actin holophosphatase that terminates signaling in the mammalian integrated stress response (ISR) in the pre-dephosphorylation complex with its substrate, translation initiation factor 2α (eIF2α). G-actin, whose essential role in eIF2α dephosphorylation is supported crystallographically, biochemically and genetically, aligns the catalytic and regulatory subunits, creating a composite surface that engages the N-terminal domain of eIF2α to position the distant phosphoserine-51 at the active site. Substrate residues that mediate affinity for the holophosphatase also make critical contacts with eIF2α kinases. Thus, a convergent process of higher-order substrate recognition specifies functionally antagonistic phosphorylation and dephosphorylation in the ISR.
Organizational Affiliation:
Cambridge Institute for Medical Research, University of Cambridge, Cambridge, UK.