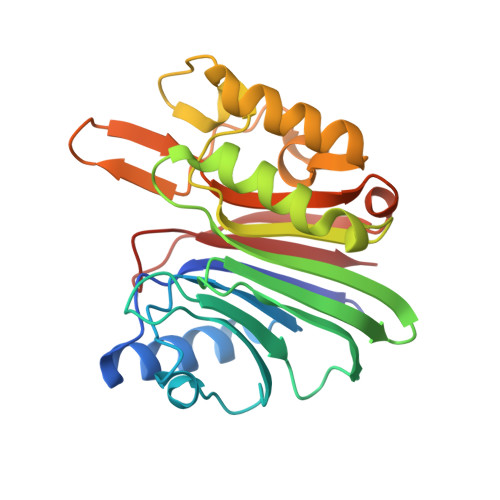
Structural dissection of sequence recognition and catalytic mechanism of human LINE-1 endonuclease.
Miller, I., Totrov, M., Korotchkina, L., Kazyulkin, D.N., Gudkov, A.V., Korolev, S.(2021) Nucleic Acids Res 49: 11350-11366
- PubMed: 34554261
- DOI: https://doi.org/10.1093/nar/gkab826
- Primary Citation of Related Structures:
7N8K, 7N8S, 7N94 - PubMed Abstract:
Long interspersed nuclear element-1 (L1) is an autonomous non-LTR retrotransposon comprising ∼20% of the human genome. L1 self-propagation causes genomic instability and is strongly associated with aging, cancer and other diseases. The endonuclease domain of L1's ORFp2 protein (L1-EN) initiates de novo L1 integration by nicking the consensus sequence 5'-TTTTT/AA-3'. In contrast, related nucleases including structurally conserved apurinic/apyrimidinic endonuclease 1 (APE1) are non-sequence specific. To investigate mechanisms underlying sequence recognition and catalysis by L1-EN, we solved crystal structures of L1-EN complexed with DNA substrates. This showed that conformational properties of the preferred sequence drive L1-EN's sequence-specificity and catalysis. Unlike APE1, L1-EN does not bend the DNA helix, but rather causes 'compression' near the cleavage site. This provides multiple advantages for L1-EN's role in retrotransposition including facilitating use of the nicked poly-T DNA strand as a primer for reverse transcription. We also observed two alternative conformations of the scissile bond phosphate, which allowed us to model distinct conformations for a nucleophilic attack and a transition state that are likely applicable to the entire family of nucleases. This work adds to our mechanistic understanding of L1-EN and related nucleases and should facilitate development of L1-EN inhibitors as potential anticancer and antiaging therapeutics.
Organizational Affiliation:
Edward A. Doisy Department of Biochemistry and Molecular Biology, Saint Louis University School of Medicine, St. Louis, MO 63104, USA.