Significant Loop Motions in the SsoPTP Protein Tyrosine Phosphatase Allow for Dual General Acid Functionality.
Pinkston, J., Jo, J., Olsen, K.J., Comer, D., Glaittli, C.A., Loria, J.P., Johnson, S.J., Hengge, A.C.(2021) Biochemistry 60: 2888-2901
- PubMed: 34496202
- DOI: https://doi.org/10.1021/acs.biochem.1c00365
- Primary Citation of Related Structures:
7MPC, 7MPD - PubMed Abstract:
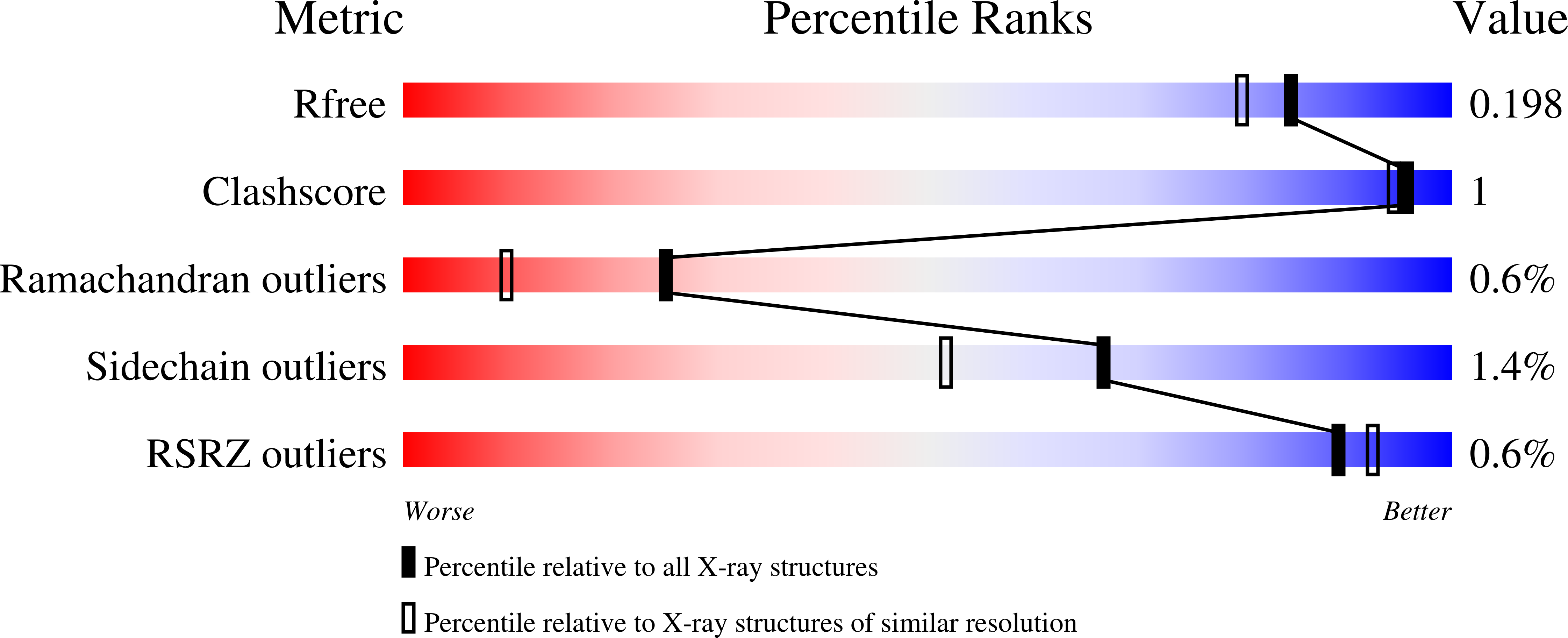
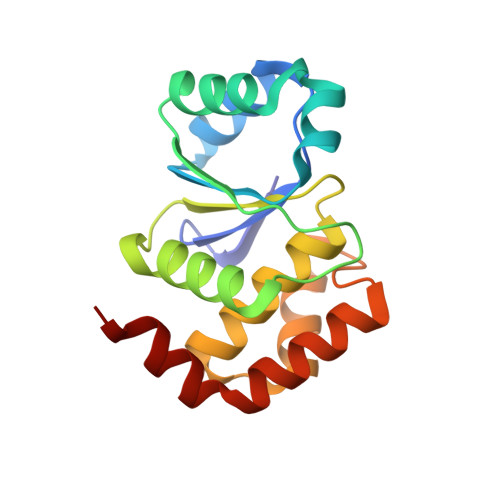
Conformational dynamics are important factors in the function of enzymes, including protein tyrosine phosphatases (PTPs). Crystal structures of PTPs first revealed the motion of a protein loop bearing a conserved catalytic aspartic acid, and subsequent nuclear magnetic resonance and computational analyses have shown the presence of motions, involved in catalysis and allostery, within and beyond the active site. The tyrosine phosphatase from the thermophilic and acidophilic Sulfolobus solfataricus (SsoPTP) displays motions of its acid loop together with dynamics of its phosphoryl-binding P-loop and the Q-loop, the first instance of such motions in a PTP. All three loops share the same exchange rate, implying their motions are coupled. Further evidence of conformational flexibility comes from mutagenesis, kinetics, and isotope effect data showing that E40 can function as an alternate general acid to protonate the leaving group when the conserved acid, D69, is mutated to asparagine. SsoPTP is not the first PTP to exhibit an alternate general acid (after VHZ and TkPTP), but E40 does not correspond to the sequence or structural location of the alternate general acids in those precedents. A high-resolution X-ray structure with the transition state analogue vanadate clarifies the role of the active site arginine R102, which varied in structures of substrates bound to a catalytically inactive mutant. The coordinated motions of all three functional loops in SsoPTP, together with the function of an alternate general acid, suggest that catalytically competent conformations are present in solution that have not yet been observed in crystal structures.
Organizational Affiliation:
Department of Chemistry and Biochemistry, Utah State University, Logan, Utah 84322-0300, United States.