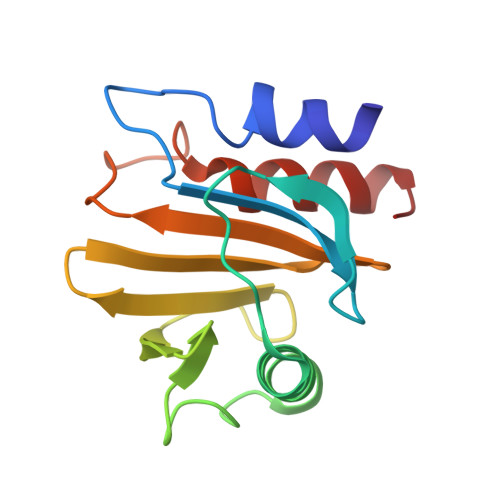
Crystal structure of timothy grass allergen Phl p 12.0101 reveals an unusual profilin dimer.
O'Malley, A., Kapingidza, A.B., Hyduke, N., Dolamore, C., Kowal, K., Chruszcz, M.(2021) Acta Biochim Pol 68: 15-22
- PubMed: 33720678
- DOI: https://doi.org/10.18388/abp.2020_5587
- Primary Citation of Related Structures:
7KYW - PubMed Abstract:
Timothy grass pollen is a source of potent allergens. Among them, Phl p 1 and Phl p 5 are thought to be the most important, as a majority of timothy grass-allergic individuals have IgE antibodies directed against these two allergens. The profilin from timothy grass (Phl p 12) has been registered as a minor allergen, with up to 35% of individuals in populations of grass pollen allergic patients showing IgE binding to Phl p 12. Profilins are primarily minor allergens and are known for a high likelihood of co-sensitization as well as cross-reactivity situations caused by their sequence and structure similarity. The crystal structure of Phl p 12.0101 was determined and it revealed that this allergen may form an unusual dimer not previously observed among any profilins. For example, the Phl p 12 dimer has a completely different geometry and interface when compared with the latex profilin (Hev b 8) dimer that has its crystal structure determined. The structure of Phl p 12.0101 is described in the context of allergenic sensitization and allergy diagnostics. Moreover, the structure of the Phl p 12.0101 dimer is discussed, taking into account the production of recombinant allergens and their storage.
Organizational Affiliation:
Department of Chemistry and Biochemistry, University of South Carolina, Columbia, SC 29208, USA.