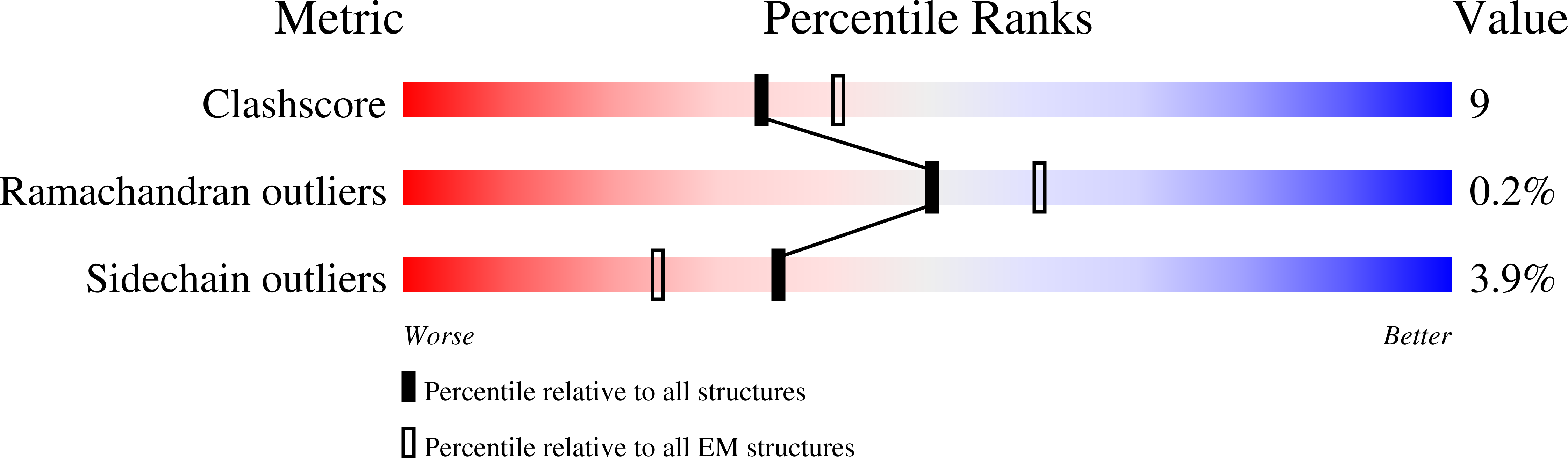Structural Basis of Zika Virus Specific Neutralization in Subsequent Flavivirus Infections.
Sevvana, M., Rogers, T.F., Miller, A.S., Long, F., Klose, T., Beutler, N., Lai, Y.C., Parren, M., Walker, L.M., Buda, G., Burton, D.R., Rossmann, M.G., Kuhn, R.J.(2020) Viruses 12
- PubMed: 33255202
- DOI: https://doi.org/10.3390/v12121346
- Primary Citation of Related Structures:
7KCR - PubMed Abstract:
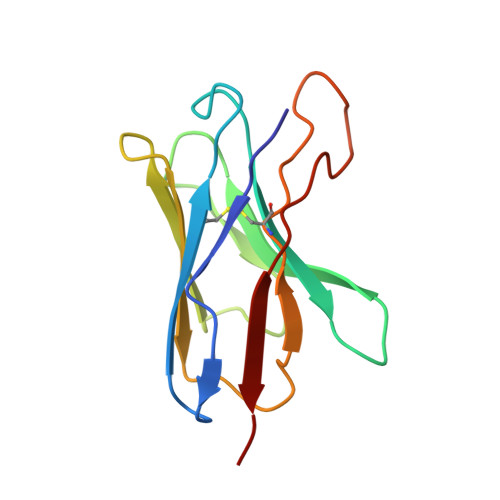
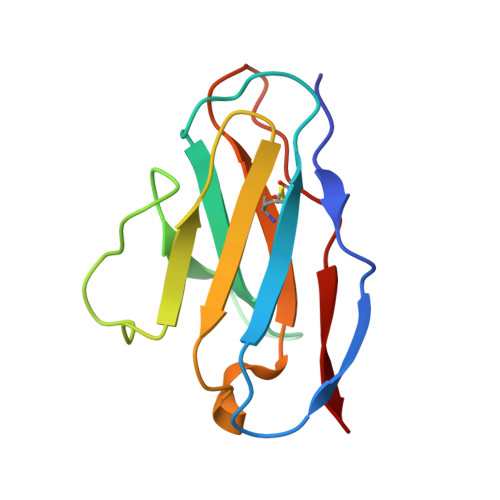
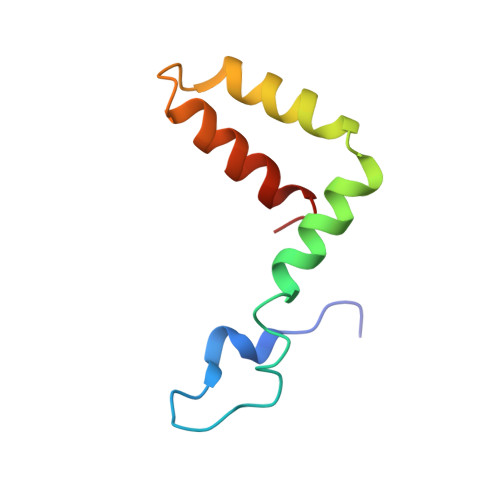
Zika virus (ZIKV), a mosquito-borne human flavivirus that causes microcephaly and other neurological disorders, has been a recent focus for the development of flavivirus vaccines and therapeutics. We report here a 4.0 Å resolution structure of the mature ZIKV in complex with ADI-30056, a ZIKV-specific human monoclonal antibody (hMAb) isolated from a ZIKV infected donor with a prior dengue virus infection. The structure shows that the hMAb interactions span across the E protein dimers on the virus surface, inhibiting conformational changes required for the formation of infectious fusogenic trimers similar to the hMAb, ZIKV-117. Structure-based functional analysis, and structure and sequence comparisons, identified ZIKV residues essential for neutralization and crucial for the evolution of highly potent E protein crosslinking Abs in ZIKV. Thus, this epitope, ZIKV's "Achilles heel", defined by the contacts between ZIKV and ADI-30056, could be a suitable target for the design of therapeutic antibodies.
Organizational Affiliation:
Department of Biological Sciences, Purdue University, West Lafayette, IN 47907, USA.