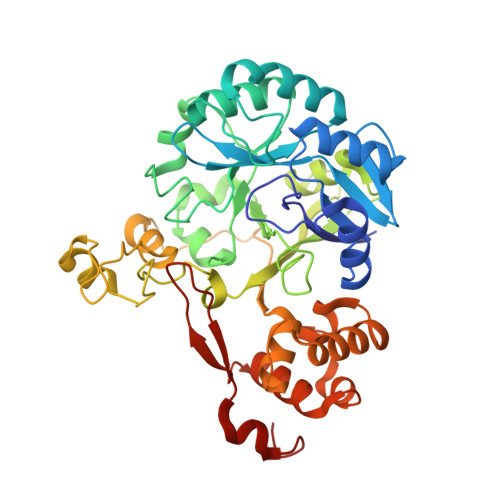
Structural basis for recognition of distinct deaminated DNA lesions by endonuclease Q.
Shi, K., Moeller, N.H., Banerjee, S., McCann, J.L., Carpenter, M.A., Yin, L., Moorthy, R., Orellana, K., Harki, D.A., Harris, R.S., Aihara, H.(2021) Proc Natl Acad Sci U S A 118
- PubMed: 33658373
- DOI: https://doi.org/10.1073/pnas.2021120118
- Primary Citation of Related Structures:
7K30, 7K31, 7K32, 7K33 - PubMed Abstract:
Spontaneous deamination of DNA cytosine and adenine into uracil and hypoxanthine, respectively, causes C to T and A to G transition mutations if left unrepaired. Endonuclease Q (EndoQ) initiates the repair of these premutagenic DNA lesions in prokaryotes by cleaving the phosphodiester backbone 5' of either uracil or hypoxanthine bases or an apurinic/apyrimidinic (AP) lesion generated by the excision of these damaged bases. To understand how EndoQ achieves selectivity toward these structurally diverse substrates without cleaving undamaged DNA, we determined the crystal structures of Pyrococcus furiosus EndoQ bound to DNA substrates containing uracil, hypoxanthine, or an AP lesion. The structures show that substrate engagement by EndoQ depends both on a highly distorted conformation of the DNA backbone, in which the target nucleotide is extruded out of the helix, and direct hydrogen bonds with the deaminated bases. A concerted swing motion of the zinc-binding and C-terminal helical domains of EndoQ toward its catalytic domain allows the enzyme to clamp down on a sharply bent DNA substrate, shaping a deep active-site pocket that accommodates the extruded deaminated base. Within this pocket, uracil and hypoxanthine bases interact with distinct sets of amino acid residues, with positioning mediated by an essential magnesium ion. The EndoQ-DNA complex structures reveal a unique mode of damaged DNA recognition and provide mechanistic insights into the initial step of DNA damage repair by the alternative excision repair pathway. Furthermore, we demonstrate that the unique activity of EndoQ is useful for studying DNA deamination and repair in mammalian systems.
Organizational Affiliation:
Department of Biochemistry, Molecular Biology, and Biophysics, University of Minnesota, Minneapolis, MN 55455.