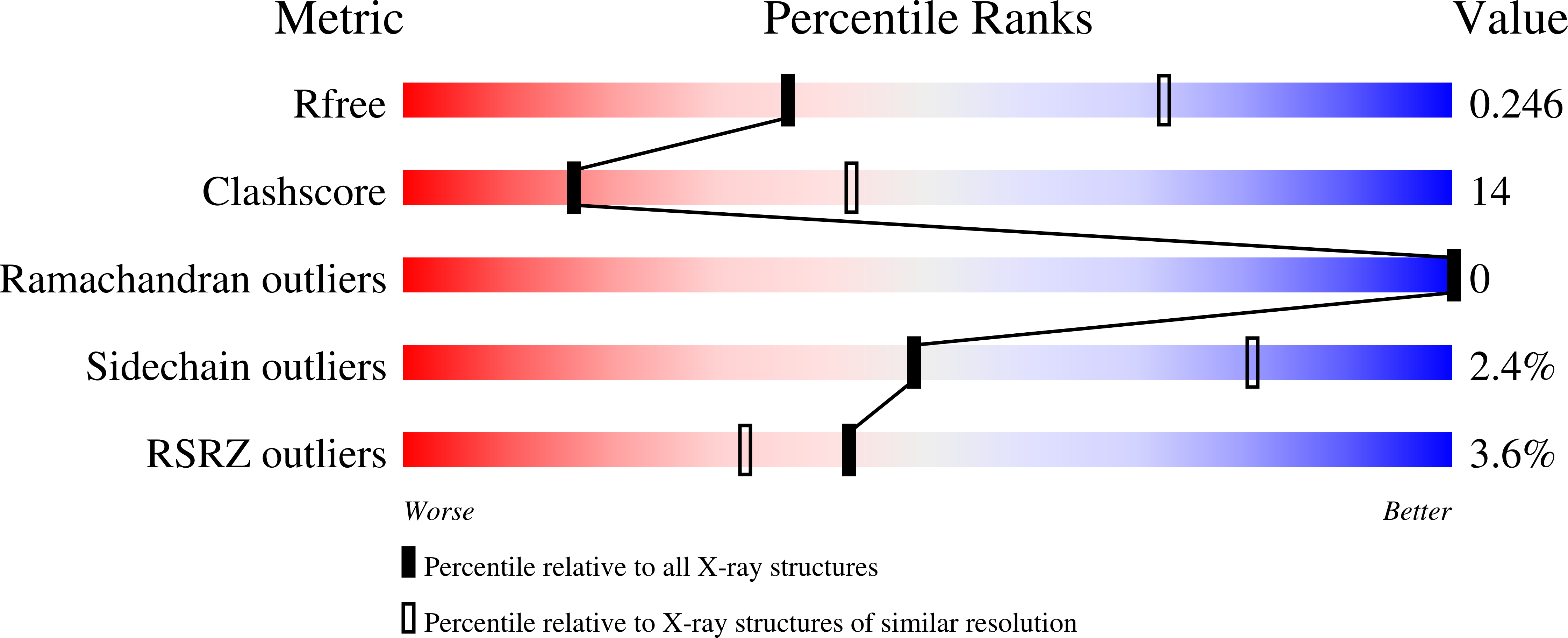
The novel anti-CRISPR AcrIIA22 relieves DNA torsion in target plasmids and impairs SpyCas9 activity.
Forsberg, K.J., Schmidtke, D.T., Werther, R., Uribe, R.V., Hausman, D., Sommer, M.O.A., Stoddard, B.L., Kaiser, B.K., Malik, H.S.(2021) PLoS Biol 19: e3001428-e3001428
- PubMed: 34644300
- DOI: https://doi.org/10.1371/journal.pbio.3001428
- Primary Citation of Related Structures:
7JTA - PubMed Abstract:
To overcome CRISPR-Cas defense systems, many phages and mobile genetic elements (MGEs) encode CRISPR-Cas inhibitors called anti-CRISPRs (Acrs). Nearly all characterized Acrs directly bind Cas proteins to inactivate CRISPR immunity. Here, using functional metagenomic selection, we describe AcrIIA22, an unconventional Acr found in hypervariable genomic regions of clostridial bacteria and their prophages from human gut microbiomes. AcrIIA22 does not bind strongly to SpyCas9 but nonetheless potently inhibits its activity against plasmids. To gain insight into its mechanism, we obtained an X-ray crystal structure of AcrIIA22, which revealed homology to PC4-like nucleic acid-binding proteins. Based on mutational analyses and functional assays, we deduced that acrIIA22 encodes a DNA nickase that relieves torsional stress in supercoiled plasmids. This may render them less susceptible to SpyCas9, which uses free energy from negative supercoils to form stable R-loops. Modifying DNA topology may provide an additional route to CRISPR-Cas resistance in phages and MGEs.
Organizational Affiliation:
Division of Basic Sciences, Fred Hutchinson Cancer Research Center, Seattle, Washington, United States of America.