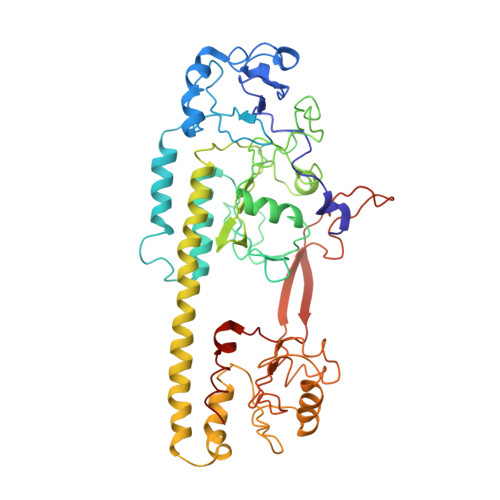
High-resolution crystal structures of transient intermediates in the phytochrome photocycle.
Carrillo, M., Pandey, S., Sanchez, J., Noda, M., Poudyal, I., Aldama, L., Malla, T.N., Claesson, E., Wahlgren, W.Y., Feliz, D., Srajer, V., Maj, M., Castillon, L., Iwata, S., Nango, E., Tanaka, R., Tanaka, T., Fangjia, L., Tono, K., Owada, S., Westenhoff, S., Stojkovic, E.A., Schmidt, M.(2021) Structure 29: 743
- PubMed: 33756101
- DOI: https://doi.org/10.1016/j.str.2021.03.004
- Primary Citation of Related Structures:
7JR5, 7JRI - PubMed Abstract:
Phytochromes are red/far-red light photoreceptors in bacteria to plants, which elicit a variety of important physiological responses. They display a reversible photocycle between the resting Pr state and the light-activated Pfr state. Light signals are transduced as structural change through the entire protein to modulate its activity. It is unknown how the Pr-to-Pfr interconversion occurs, as the structure of intermediates remains notoriously elusive. Here, we present short-lived crystal structures of the photosensory core modules of the bacteriophytochrome from myxobacterium Stigmatella aurantiaca captured by an X-ray free electron laser 5 ns and 33 ms after light illumination of the Pr state. We observe large structural displacements of the covalently bound bilin chromophore, which trigger a bifurcated signaling pathway that extends through the entire protein. The snapshots show with atomic precision how the signal progresses from the chromophore, explaining how plants, bacteria, and fungi sense red light.
Organizational Affiliation:
Department of Biology, Northeastern Illinois University, 5500 North St. Louis Avenue, Chicago, IL 60625, USA.