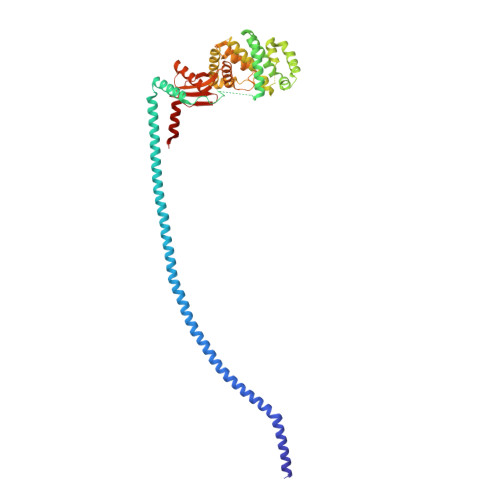
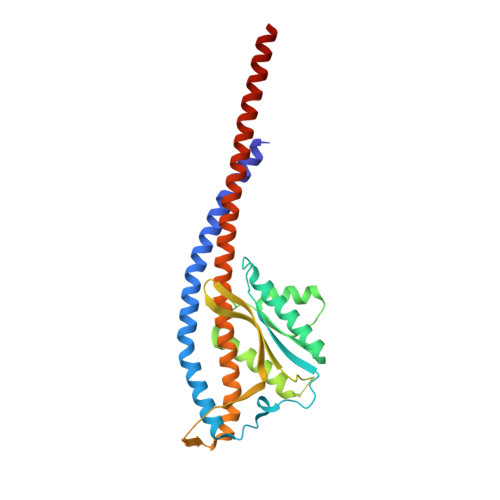
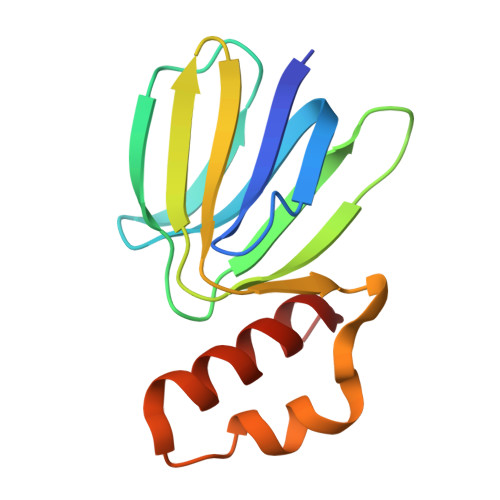
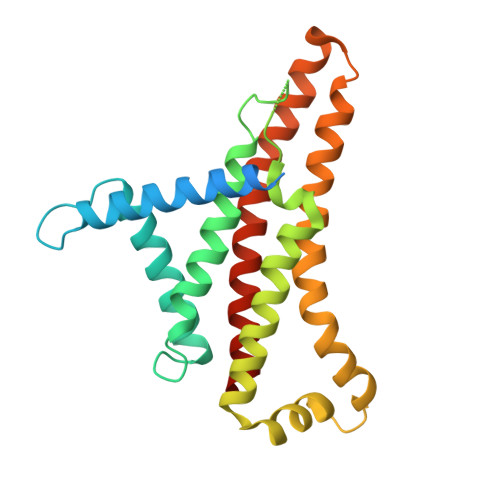
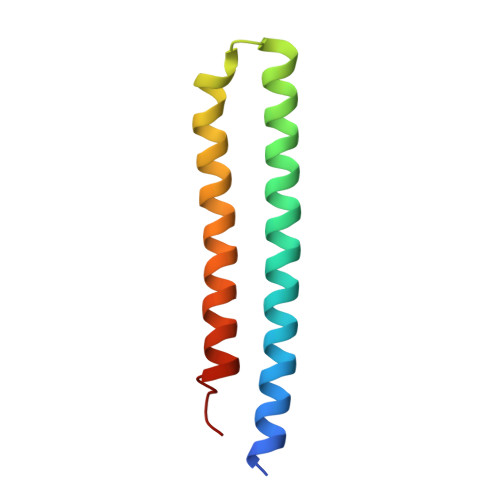
Structure of mycobacterial ATP synthase bound to the tuberculosis drug bedaquiline.
Guo, H., Courbon, G.M., Bueler, S.A., Mai, J., Liu, J., Rubinstein, J.L.(2021) Nature 589: 143-147
- PubMed: 33299175
- DOI: https://doi.org/10.1038/s41586-020-3004-3
- Primary Citation of Related Structures:
7JG5, 7JG6, 7JG7, 7JG8, 7JG9, 7JGA, 7JGB, 7JGC - PubMed Abstract:
Tuberculosis-the world's leading cause of death by infectious disease-is increasingly resistant to current first-line antibiotics 1 . The bacterium Mycobacterium tuberculosis (which causes tuberculosis) can survive low-energy conditions, allowing infections to remain dormant and decreasing their susceptibility to many antibiotics 2 . Bedaquiline was developed in 2005 from a lead compound identified in a phenotypic screen against Mycobacterium smegmatis 3 . This drug can sterilize even latent M. tuberculosis infections 4 and has become a cornerstone of treatment for multidrug-resistant and extensively drug-resistant tuberculosis 1,5,6 . Bedaquiline targets the mycobacterial ATP synthase 3 , which is an essential enzyme in the obligate aerobic Mycobacterium genus 3,7 , but how it binds the intact enzyme is unknown. Here we determined cryo-electron microscopy structures of M. smegmatis ATP synthase alone and in complex with bedaquiline. The drug-free structure suggests that hook-like extensions from the α-subunits prevent the enzyme from running in reverse, inhibiting ATP hydrolysis and preserving energy in hypoxic conditions. Bedaquiline binding induces large conformational changes in the ATP synthase, creating tight binding pockets at the interface of subunits a and c that explain the potency of this drug as an antibiotic for tuberculosis.
Organizational Affiliation:
Molecular Medicine Program, The Hospital for Sick Children, Toronto, Ontario, Canada.