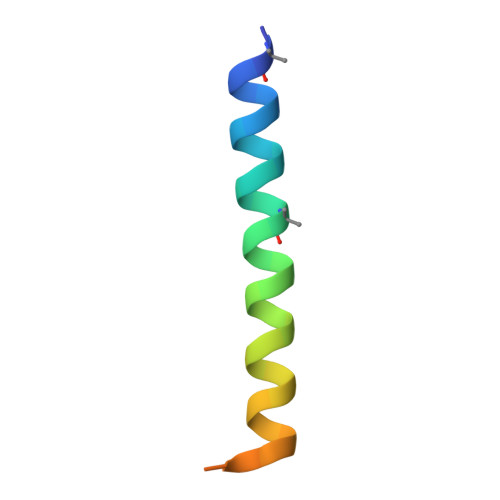
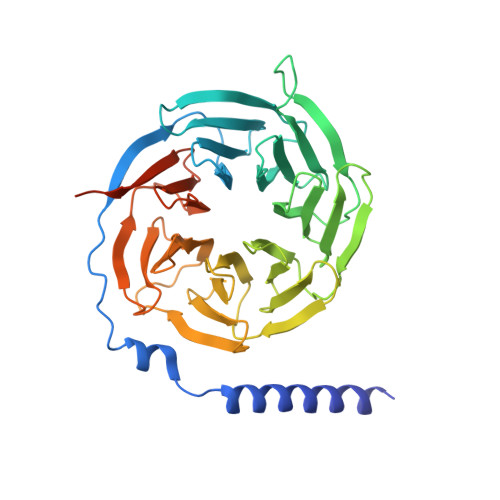
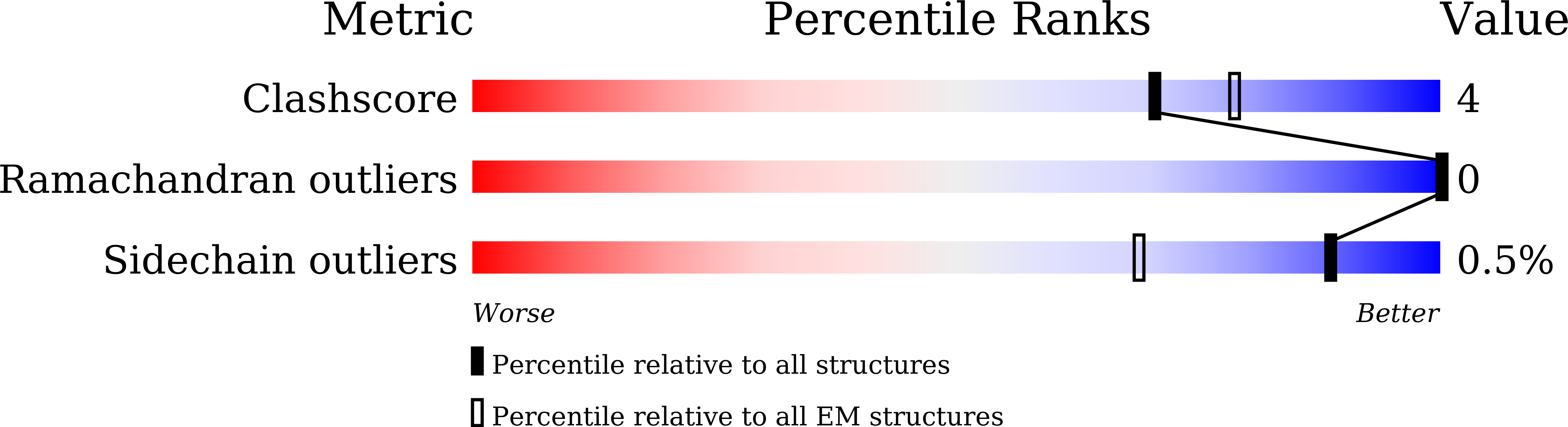Structural insights into multiplexed pharmacological actions of tirzepatide and peptide 20 at the GIP, GLP-1 or glucagon receptors.
Zhao, F., Zhou, Q., Cong, Z., Hang, K., Zou, X., Zhang, C., Chen, Y., Dai, A., Liang, A., Ming, Q., Wang, M., Chen, L.N., Xu, P., Chang, R., Feng, W., Xia, T., Zhang, Y., Wu, B., Yang, D., Zhao, L., Xu, H.E., Wang, M.W.(2022) Nat Commun 13: 1057-1057
- PubMed: 35217653
- DOI: https://doi.org/10.1038/s41467-022-28683-0
- Primary Citation of Related Structures:
7FIM, 7FIN, 7FIY, 7V35, 7VAB, 7VBH, 7VBI - PubMed Abstract:
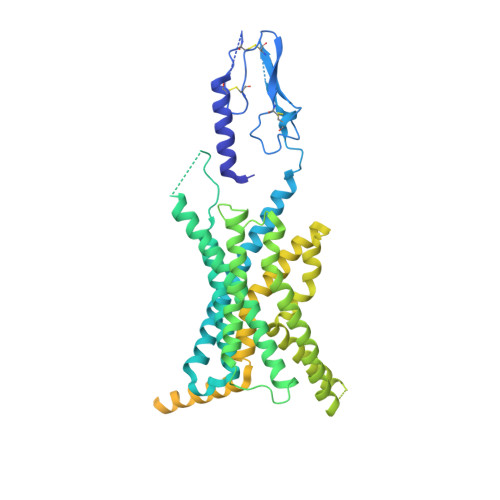
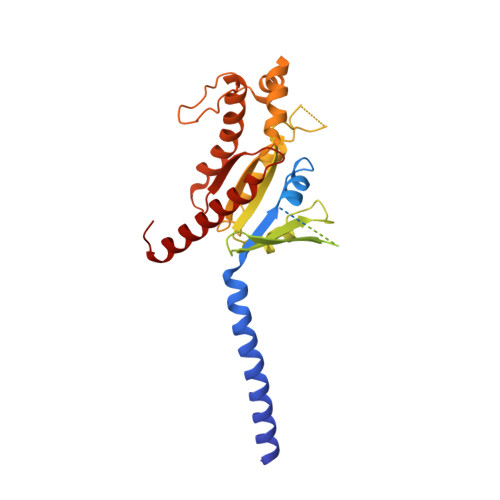
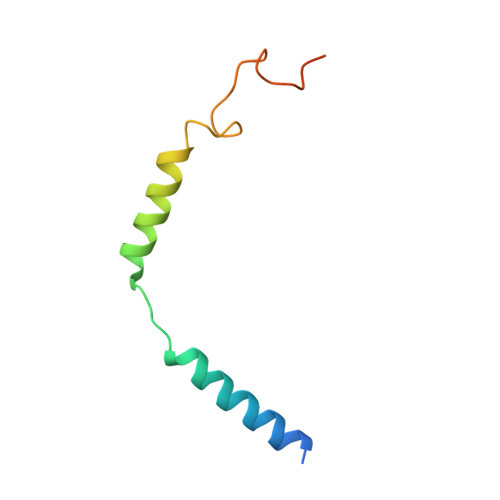
Glucose homeostasis, regulated by glucose-dependent insulinotropic polypeptide (GIP), glucagon-like peptide-1 (GLP-1) and glucagon (GCG) is critical to human health. Several multi-targeting agonists at GIPR, GLP-1R or GCGR, developed to maximize metabolic benefits with reduced side-effects, are in clinical trials to treat type 2 diabetes and obesity. To elucidate the molecular mechanisms by which tirzepatide, a GIPR/GLP-1R dual agonist, and peptide 20, a GIPR/GLP-1R/GCGR triagonist, manifest their multiplexed pharmacological actions over monoagonists such as semaglutide, we determine cryo-electron microscopy structures of tirzepatide-bound GIPR and GLP-1R as well as peptide 20-bound GIPR, GLP-1R and GCGR. The structures reveal both common and unique features for the dual and triple agonism by illustrating key interactions of clinical relevance at the near-atomic level. Retention of glucagon function is required to achieve such an advantage over GLP-1 monotherapy. Our findings provide valuable insights into the structural basis of functional versatility of tirzepatide and peptide 20.
Organizational Affiliation:
School of Pharmacy, Fudan University, Shanghai, China.