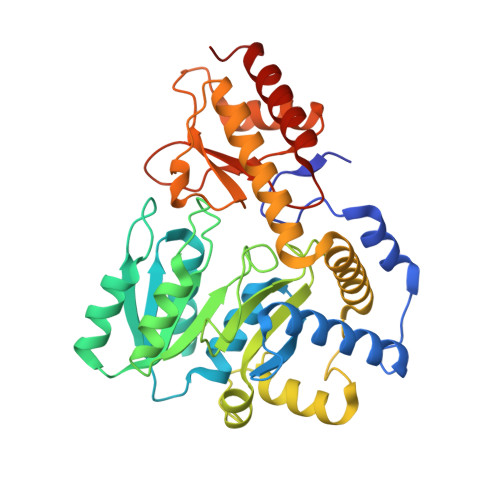
Structural characterization of a 2-aminoethylphosphonate:pyruvate aminotransferase from Pseudomonas aeruginosa PAO1.
Jia, H., Chen, Y., Chen, Y., Liu, R., Zhang, Q., Bartlam, M.(2021) Biochem Biophys Res Commun 552: 114-119
- PubMed: 33743347
- DOI: https://doi.org/10.1016/j.bbrc.2021.03.046
- Primary Citation of Related Structures:
7E7G - PubMed Abstract:
2-aminoethylphosphonate:pyruvate aminotransferase (AEPT) is a pyridoxal 5'-phosphate (PLP)-dependent enzyme that mediates the first step in the AEP degradation pathway. It catalyzes the transamination of 2-aminoethylphosphonate (AEP) with pyruvate to phosphonoacetaldehyde and l-alanine respectively. Although the enzyme is widely present in microorganisms, there are few reports on the structure and function of AEPT to date. Here we report the crystal structure of AEPT from Pseudomonas aeruginosa PAO1 (PaAEPT) to 2.35 Å resolution in the absence of the PLP cofactor. PaAEPT crystallizes in space group P2 1 2 1 2 with one monomer per asymmetric unit. Analytical ultracentrifugation analysis shows that PaAEPT forms a stable dimer in solution. Our work provides a valuable starting point for further functional and mechanistic studies of the AEP degradation pathway.
Organizational Affiliation:
State Key Laboratory of Medicinal Chemical Biology, Tianjin Key Laboratory of Protein Science and College of Life Sciences, Nankai University, Tianjin, 300071, China.