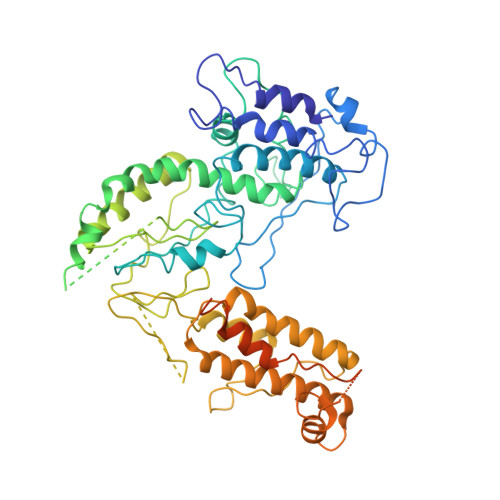
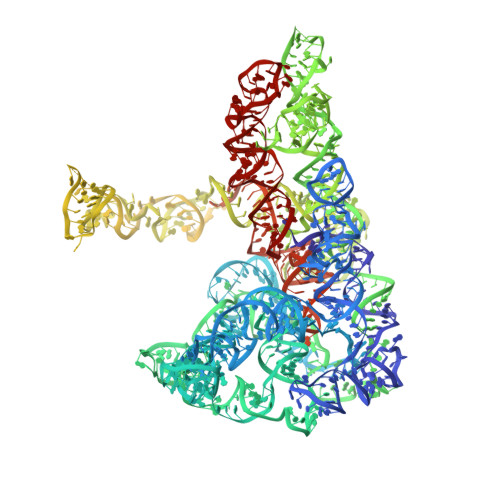
Exon and protein positioning in a pre-catalytic group II intron RNP primed for splicing.
Liu, N., Dong, X., Hu, C., Zeng, J., Wang, J., Wang, J., Wang, H.W., Belfort, M.(2020) Nucleic Acids Res 48: 11185-11198
- PubMed: 33021674
- DOI: https://doi.org/10.1093/nar/gkaa773
- Primary Citation of Related Structures:
7D0F, 7D0G, 7D1A - PubMed Abstract:
Group II introns are the putative progenitors of nuclear spliceosomal introns and use the same two-step splicing pathway. In the cell, the intron RNA forms a ribonucleoprotein (RNP) complex with the intron-encoded protein (IEP), which is essential for splicing. Although structures of spliced group II intron RNAs and RNP complexes have been characterized, structural insights into the splicing process remain enigmatic due to lack of pre-catalytic structural models. Here, we report two cryo-EM structures of endogenously produced group II intron RNPs trapped in their pre-catalytic state. Comparison of the catalytically activated precursor RNP to its previously reported spliced counterpart allowed identification of key structural rearrangements accompanying splicing, including a remodeled active site and engagement of the exons. Importantly, altered RNA-protein interactions were observed upon splicing among the RNP complexes. Furthermore, analysis of the catalytically inert precursor RNP demonstrated the structural impact of the formation of the active site on RNP architecture. Taken together, our results not only fill a gap in understanding the structural basis of IEP-assisted group II intron splicing, but also provide parallels to evolutionarily related spliceosomal splicing.
Organizational Affiliation:
Ministry of Education Key Laboratory of Protein Sciences, Tsinghua-Peking Joint Center for Life Sciences, Beijing Advanced Innovation Center for Structural Biology, School of Life Sciences, Tsinghua University, Beijing 100084, China.