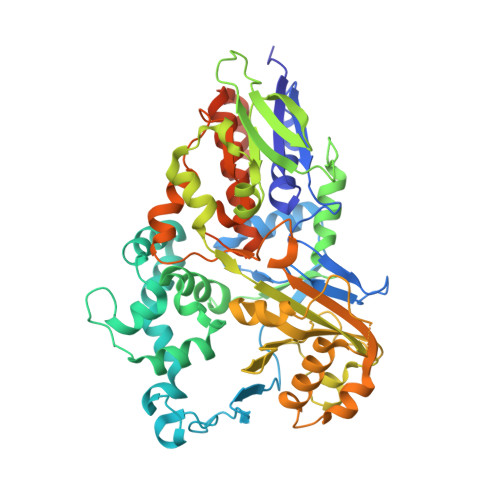
Structural basis of strict substrate recognition of l-lysine alpha-oxidase from Trichoderma viride.
Kondo, H., Kitagawa, M., Matsumoto, Y., Saito, M., Amano, M., Sugiyama, S., Tamura, T., Kusakabe, H., Inagaki, K., Imada, K.(2020) Protein Sci 29: 2213-2225
- PubMed: 32894626
- DOI: https://doi.org/10.1002/pro.3946
- Primary Citation of Related Structures:
7C3H, 7C3I, 7C3J, 7C3L - PubMed Abstract:
l-Lysine oxidase (LysOX) is a FAD-dependent homodimeric enzyme that catalyzes the oxidative deamination of l-lysine to produce α-keto-ε-aminocaproate with ammonia and hydrogen peroxide. LysOX shows strict substrate specificity for l-lysine, whereas most l-amino acid oxidases (LAAOs) exhibit broad substrate specificity for l-amino acids. Previous studies of LysOX showed that overall structural similarity to the well-studied snake venom LAAOs. However, the molecular mechanism of strict specificity for l-lysine was still unclear. We here determined the structure of LysOX in complex with l-lysine at 1.7 Å resolution. The structure revealed that the hydrogen bonding network formed by D212, D315, and A440 with two water molecules is responsible for the recognition of the side chain amino group. In addition, a narrow hole formed by five hydrophobic residues in the active site contributes to strict substrate specificity. Mutation studies demonstrated that D212 and D315 are essential for l-lysine recognition, and the D212A/D315A double mutant LysOX showed different substrate specificity from LysOX. Moreover, the structural basis of the substrate specificity change has also been revealed by the structural analysis of the mutant variant and its substrate complexes. These results clearly explain the molecular mechanism of the strict specificity of LysOX and suggest that LysOX is a potential candidate for a template to design LAAOs specific to other l-amino acids.
Organizational Affiliation:
Department of Macromolecular Science, Graduate School of Science, Osaka University, Osaka, Japan.