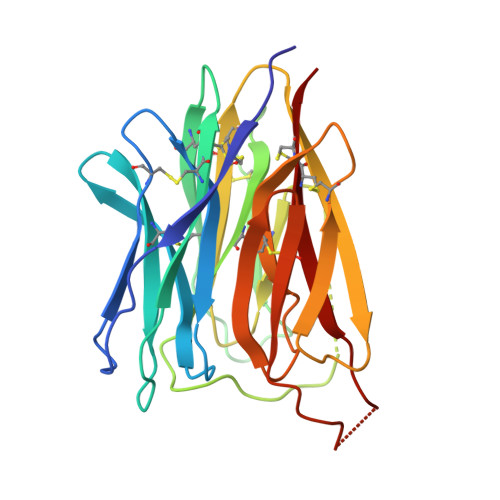
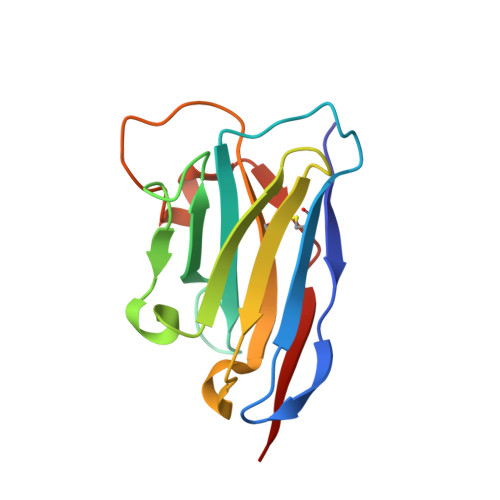
Molecular interactions of PCSK9 with an inhibitory nanobody, CAP1 and HLA-C: Functional regulation of LDLR levels.
Fruchart Gaillard, C., Ouadda, A.B.D., Ciccone, L., Girard, E., Mikaeeli, S., Evagelidis, A., Le Devehat, M., Susan-Resiga, D., Lajeunesse, E.C., Nozach, H., Ramos, O.H.P., Thureau, A., Legrand, P., Prat, A., Dive, V., Seidah, N.G.(2022) Mol Metab 67: 101662-101662
- PubMed: 36566984
- DOI: https://doi.org/10.1016/j.molmet.2022.101662
- Primary Citation of Related Structures:
7ANQ - PubMed Abstract:
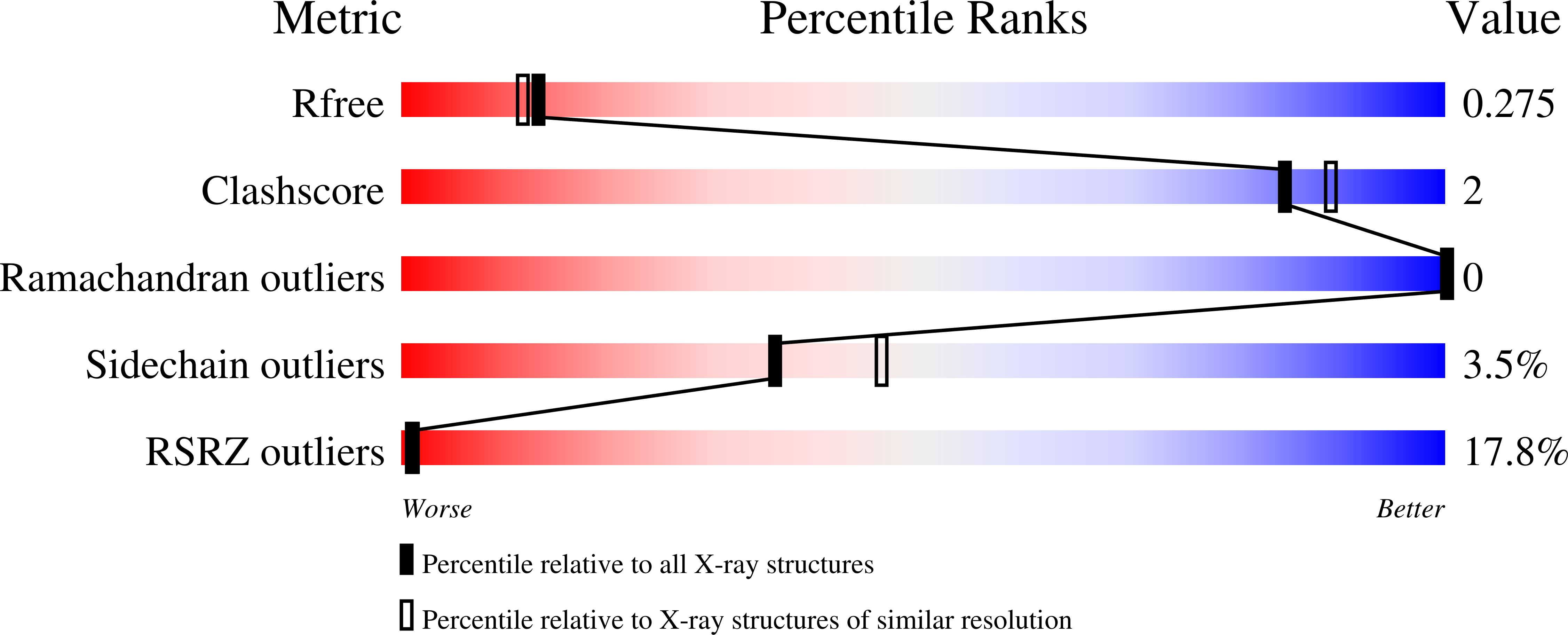
The liver-derived circulating PCSK9 enhances the degradation of the LDL receptor (LDLR) in endosomes/lysosomes. PCSK9 inhibition or silencing is presently used in clinics worldwide to reduce LDL-cholesterol, resulting in lower incidence of cardiovascular disease and possibly cancer/metastasis. The mechanism by which the PCSK9-LDLR complex is sorted to degradation compartments is not fully understood. We previously suggested that out of the three M1, M2 and M3 subdomains of the C-terminal Cys/His-rich-domain (CHRD) of PCSK9, only M2 is critical for the activity of extracellular of PCSK9 on cell surface LDLR. This likely implicates the binding of M2 to an unknown membrane-associated "protein X" that would escort the complex to endosomes/lysosomes for degradation. We reported that a nanobody P1.40 binds the M1 and M3 domains of the CHRD and inhibits the function of PCSK9. It was also reported that the cytosolic adenylyl cyclase-associated protein 1 (CAP1) could bind M1 and M3 subdomains and enhance the activity of PCSK9. In this study, we determined the 3-dimensional structure of the CHRD-P1.40 complex to understand the intricate interplay between P1.40, CAP1 and PCSK9 and how they regulate LDLR degradation.
Organizational Affiliation:
Université Paris-Saclay, CEA, INRAE, Département Médicaments et Technologies pour la Santé (DMTS), SIMoS, 91191 Gif-sur-Yvette, France.