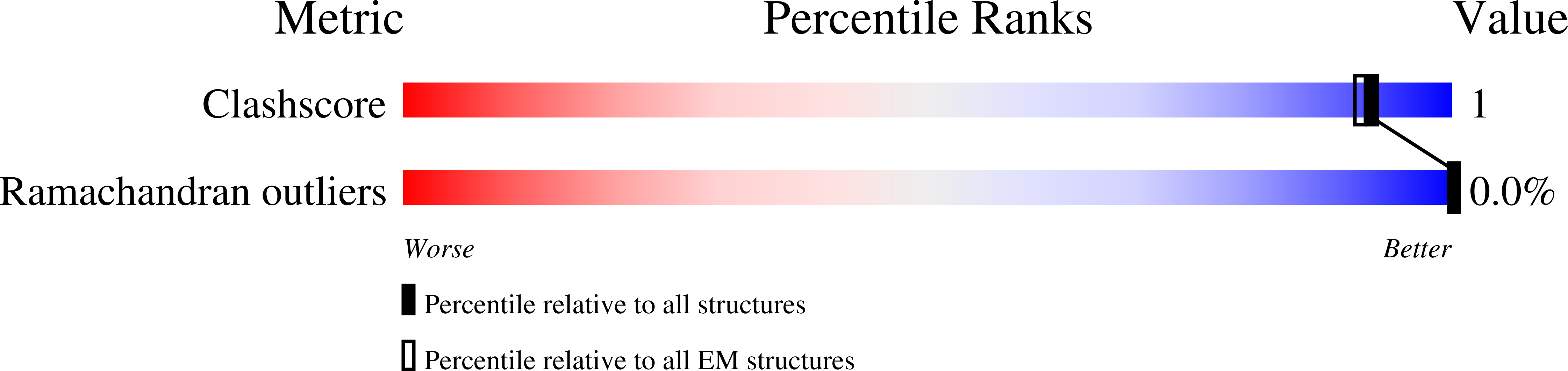Interface mobility between monomers in dimeric bovine ATP synthase participates in the ultrastructure of inner mitochondrial membranes.
Spikes, T.E., Montgomery, M.G., Walker, J.E.(2021) Proc Natl Acad Sci U S A 118
- PubMed: 33542155
- DOI: https://doi.org/10.1073/pnas.2021012118
- Primary Citation of Related Structures:
7AJB, 7AJC, 7AJD, 7AJE, 7AJF, 7AJG, 7AJH, 7AJI, 7AJJ - PubMed Abstract:
The ATP synthase complexes in mitochondria make the ATP required to sustain life by a rotary mechanism. Their membrane domains are embedded in the inner membranes of the organelle, and they dimerize via interactions between their membrane domains. The dimers form extensive chains along the tips of the cristae with the two rows of monomeric catalytic domains extending into the mitochondrial matrix at an angle to each other. Disruption of the interface between dimers by mutation affects the morphology of the cristae severely. By analysis of particles of purified dimeric bovine ATP synthase by cryo-electron microscopy, we have shown that the angle between the central rotatory axes of the monomeric complexes varies between ca. 76 and 95°. These particles represent active dimeric ATP synthase. Some angular variations arise directly from the catalytic mechanism of the enzyme, and others are independent of catalysis. The monomer-monomer interaction is mediated mainly by j subunits attached to the surface of wedge-shaped protein-lipid structures in the membrane domain of the complex, and the angular variation arises from rotational and translational changes in this interaction, and combinations of both. The structures also suggest how the dimeric ATP synthases might be interacting with each other to form the characteristic rows along the tips of the cristae via other interwedge contacts, molding themselves to the range of oligomeric arrangements observed by tomography of mitochondrial membranes, and at the same time allowing the ATP synthase to operate under the range of physiological conditions that influence the structure of the cristae.
Organizational Affiliation:
The Medical Research Council Mitochondrial Biology Unit, University of Cambridge, Cambridge, CB2 0XY, United Kingdom.