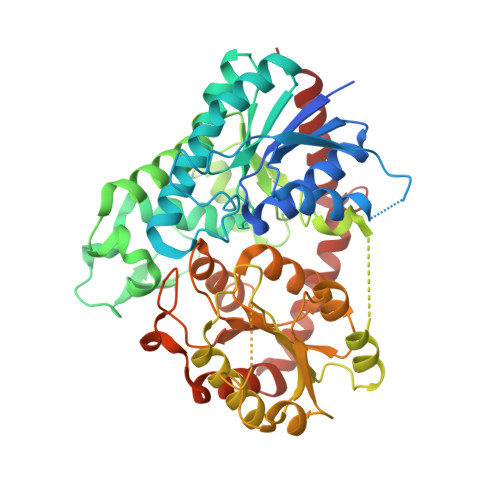
Structural basis for substrate recognition in the Phytolacca americana glycosyltransferase PaGT3.
Maharjan, R., Fukuda, Y., Nakayama, T., Nakayama, T., Hamada, H., Ozaki, S.I., Inoue, T.(2022) Acta Crystallogr D Struct Biol 78: 379-389
- PubMed: 35234151
- DOI: https://doi.org/10.1107/S2059798322000869
- Primary Citation of Related Structures:
7VEJ, 7VEK, 7VEL - PubMed Abstract:
Capsaicinoids are phenolic compounds that have health benefits. However, the pungency and poor water solubility of these compounds limit their exploitation. Glycosylation is a powerful method to improve water solubility and reduce pungency while preserving bioactivity. PaGT3, a uridine diphosphate glycosyltransferase (UGT) from Phytolacca americana, is known for its ability to glycosylate capsaicinoids and other phenolic compounds. While structural information on several UGTs is available, structures of UGTs that can glycosylate a range of phenolic compounds are rare. To fill this gap, crystal structures of PaGT3 with a sugar-donor analogue (UDP-2-fluoroglucose) and the acceptors capsaicin and kaempferol were determined. PaGT3 adopts a GT-B-fold structure that is highly conserved among UGTs. However, the acceptor-binding pocket in PaGT3 is hydrophobic and large, and is surrounded by longer loops. The larger acceptor-binding pocket in PaGT3 allows the enzyme to bind a range of compounds, while the flexibility of the longer loops possibly plays a role in accommodating the acceptors in the binding pocket according to their shape and size. This structural information provides insights into the acceptor-binding mechanism in UGTs that bind multiple substrates.
Organizational Affiliation:
Graduate School of Pharmaceutical Science, Osaka University, Suita, Osaka 565-0871, Japan.