Structural insights into the HIV-1 Vpr mediated ubiquitination through the Cullin-RING E3 ubiquitin ligase
Wang, D., Xu, J., Liu, Q., Xiang, Y.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| DDB1- and CUL4-associated factor 1 | 1,507 | Homo sapiens | Mutation(s): 0 Gene Names: DCAF1, KIAA0800, RIP, VPRBP EC: 2.7.11.1 | 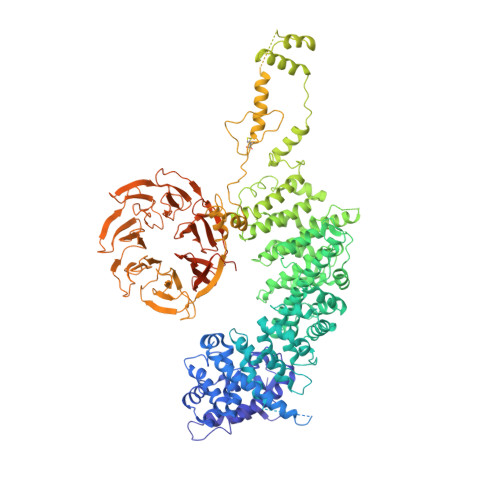 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for Q9Y4B6 (Homo sapiens) Explore Q9Y4B6 Go to UniProtKB: Q9Y4B6 | |||||
PHAROS: Q9Y4B6 GTEx: ENSG00000145041 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9Y4B6 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| DNA damage-binding protein 1 | 1,140 | Homo sapiens | Mutation(s): 0 Gene Names: DDB1, XAP1 | 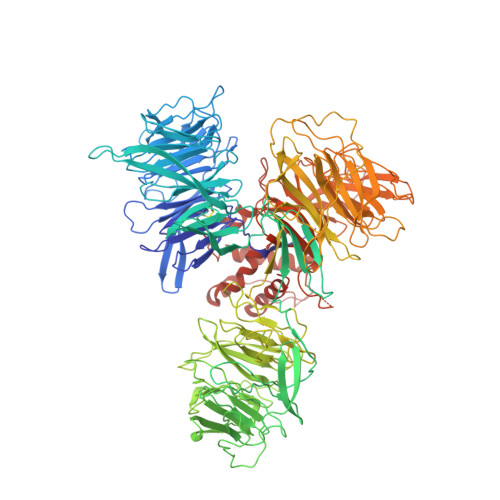 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for Q16531 (Homo sapiens) Explore Q16531 Go to UniProtKB: Q16531 | |||||
PHAROS: Q16531 GTEx: ENSG00000167986 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q16531 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Protein Vpr | 96 | Human immunodeficiency virus 1 | Mutation(s): 0 Gene Names: vpr | 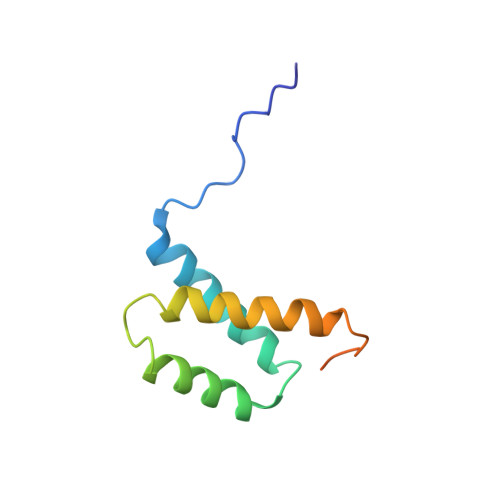 | |
UniProt | |||||
Find proteins for P12520 (Human immunodeficiency virus type 1 group M subtype B (isolate NY5)) Explore P12520 Go to UniProtKB: P12520 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P12520 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Uracil-DNA glycosylase | 220 | Homo sapiens | Mutation(s): 0 Gene Names: UNG, DGU, UNG1, UNG15 EC: 3.2.2.27 | 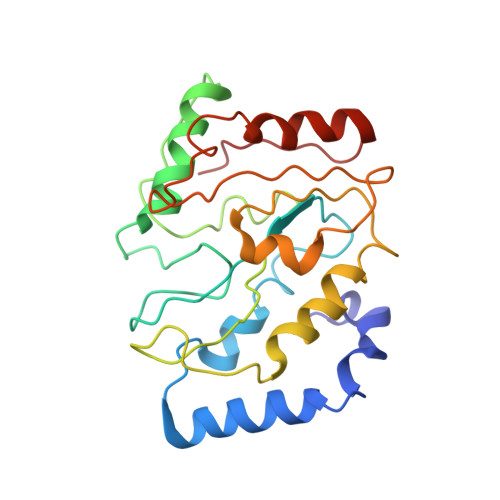 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P13051 (Homo sapiens) Explore P13051 Go to UniProtKB: P13051 | |||||
PHAROS: P13051 GTEx: ENSG00000076248 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P13051 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Task | Software Package | Version |
|---|---|---|
| RECONSTRUCTION | RELION | |
| MODEL REFINEMENT | PHENIX |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Ministry of Science and Technology (MoST, China) | China | 2016YFA0501100 |
| National Natural Science Foundation of China (NSFC) | China | 31925023 |
| National Natural Science Foundation of China (NSFC) | China | 21827810 |
| National Natural Science Foundation of China (NSFC) | China | 31861143027 |
| National Natural Science Foundation of China (NSFC) | China | 31470721 |