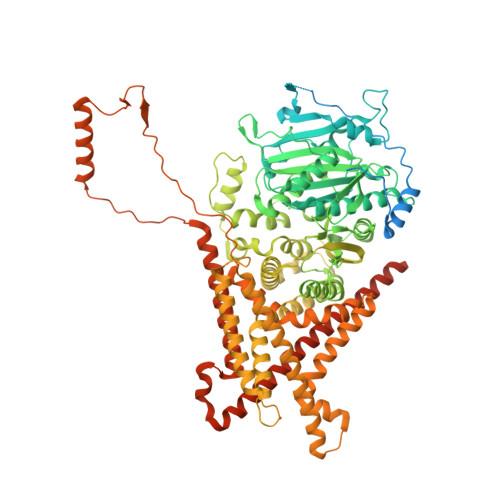
Structural basis for inhibition and regulation of a chitin synthase from Candida albicans.
Ren, Z., Chhetri, A., Guan, Z., Suo, Y., Yokoyama, K., Lee, S.Y.(2022) Nat Struct Mol Biol 29: 653-664
- PubMed: 35788183
- DOI: https://doi.org/10.1038/s41594-022-00791-x
- Primary Citation of Related Structures:
7STL, 7STM, 7STN, 7STO - PubMed Abstract:
Chitin is an essential component of the fungal cell wall. Chitin synthases (Chss) catalyze chitin formation and translocation across the membrane and are targets of antifungal agents, including nikkomycin Z and polyoxin D. Lack of structural insights into the action of these inhibitors on Chs has hampered their further development to the clinic. We present the cryo-EM structures of Chs2 from Candida albicans (CaChs2) in the apo, substrate-bound, nikkomycin Z-bound, and polyoxin D-bound states. CaChs2 adopts a unique domain-swapped dimer configuration where a conserved motif in the domain-swapped region controls enzyme activity. CaChs2 has a dual regulation mechanism where the chitin translocation tunnel is closed by the extracellular gate and plugged by a lipid molecule in the apo state to prevent non-specific leak. Analyses of substrate and inhibitor binding provide insights into the chemical logic of Chs inhibition, which can guide Chs-targeted antifungal development.
Organizational Affiliation:
Department of Biochemistry, Duke University School of Medicine, Durham, NC, USA.