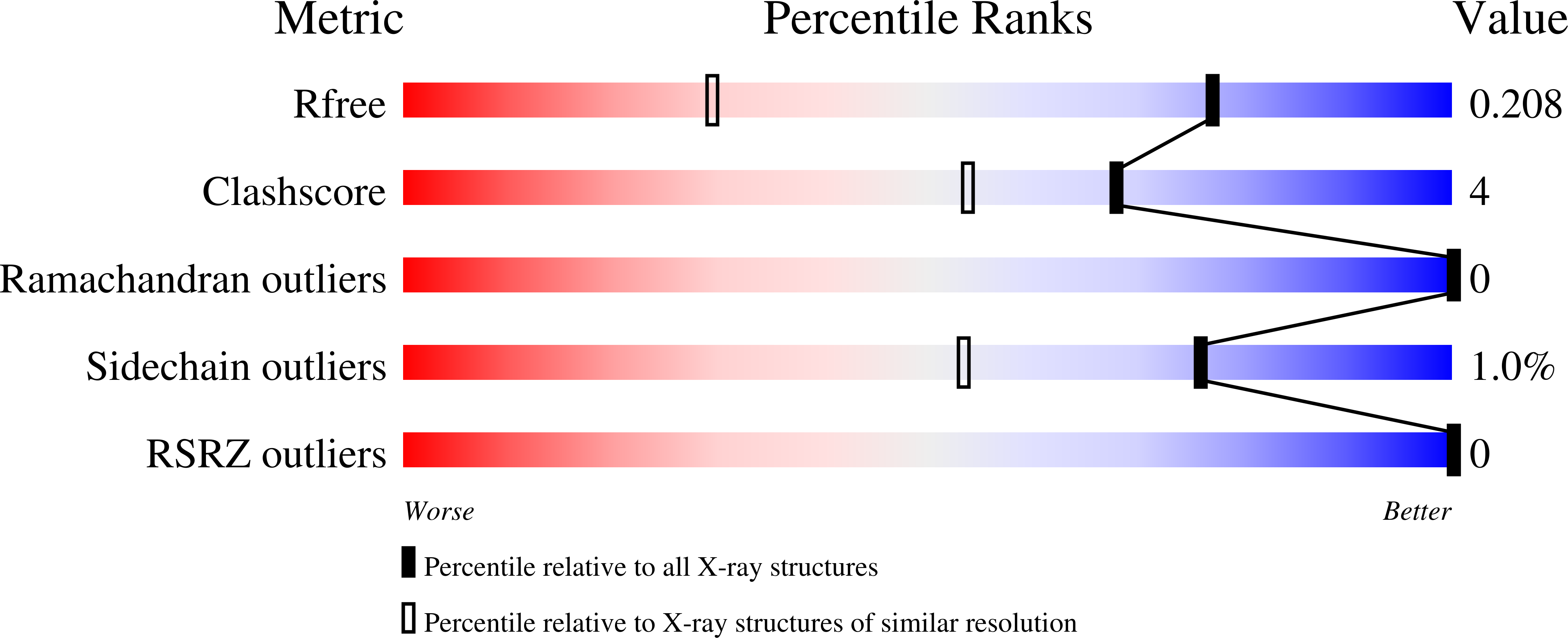
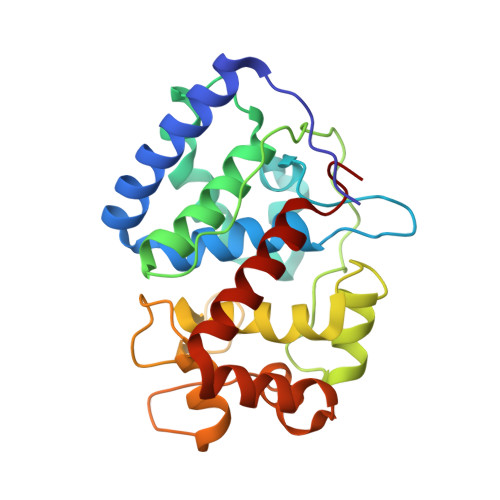
Computational analysis of the tryptophan cation radical energetics in peroxidase Compound I.
Poulos, T.L., Kim, J.S., Murarka, V.C.(2022) J Biol Inorg Chem 27: 229-237
- PubMed: 35064363
- DOI: https://doi.org/10.1007/s00775-022-01925-8
- Primary Citation of Related Structures:
7S10 - PubMed Abstract:
Three well-characterized heme peroxidases (cytochrome c peroxidase = CCP, ascorbate peroxidase = APX, and Leishmania major peroxidase = LMP) all have a Trp residue tucked under the heme stacked against the proximal His heme ligand. The reaction of peroxidases with H 2 O 2 to give Compound I results in the oxidation of this Trp to a cationic radical in CCP and LMP but not in APX. Considerable experimental data indicate that the local electrostatic environment controls whether this Trp or the porphyrin is oxidized in Compound I. Attempts have been made to place the differences between these peroxidases on a quantitative basis using computational methods. These efforts have been somewhat limited by the approximations required owing to the computational cost of using fully solvated atomistic models with well-developed forcefields. This now has changed with available GPU computing power and the associated development of software. Here we employ thermodynamic integration and multistate Bennett acceptance ratio methods to help fine-tune our understanding on the energetic differences in Trp radical stabilization in all three peroxidases. These results indicate that the local solvent structure near the redox active Trp plays a significant role in stabilization of the cationic Trp radical.
Organizational Affiliation:
Departments of Molecular Biology and Biochemistry, Chemistry, and Pharmaceutical Sciences, University of Calif, Irvine, Irvine, CA, 92697-3900, USA. poulos@uci.edu.