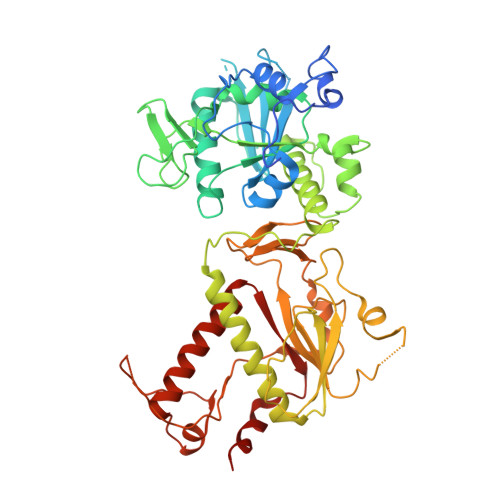
Crystal structures and fragment screening of SARS-CoV-2 NSP14 reveal details of exoribonuclease activation and mRNA capping and provide starting points for antiviral drug development.
Imprachim, N., Yosaatmadja, Y., Newman, J.A.(2023) Nucleic Acids Res 51: 475-487
- PubMed: 36546776
- DOI: https://doi.org/10.1093/nar/gkac1207
- Primary Citation of Related Structures:
7QGI, 7QIF - PubMed Abstract:
NSP14 is a dual function enzyme containing an N-terminal exonuclease domain (ExoN) and C-terminal Guanine-N7-methyltransferase (N7-MTase) domain. Both activities are essential for the viral life cycle and may be targeted for anti-viral therapeutics. NSP14 forms a complex with NSP10, and this interaction enhances the nuclease but not the methyltransferase activity. We have determined the structure of SARS-CoV-2 NSP14 in the absence of NSP10 to 1.7 Å resolution. Comparisons with NSP14/NSP10 complexes reveal significant conformational changes that occur within the NSP14 ExoN domain upon binding of NSP10, including helix to coil transitions that facilitate the formation of the ExoN active site and provide an explanation of the stimulation of nuclease activity by NSP10. We have determined the structure of NSP14 in complex with cap analogue 7MeGpppG, and observe conformational changes within a SAM/SAH interacting loop that plays a key role in viral mRNA capping offering new insights into MTase activity. We perform an X-ray fragment screen on NSP14, revealing 72 hits bound to sites of inhibition in the ExoN and MTase domains. These fragments serve as excellent starting point tools for structure guided development of NSP14 inhibitors that may be used to treat COVID-19 and potentially other future viral threats.
Organizational Affiliation:
Centre for Medicines Discovery, University of Oxford, South Parks Rd, Oxford OX1 3QU, UK.