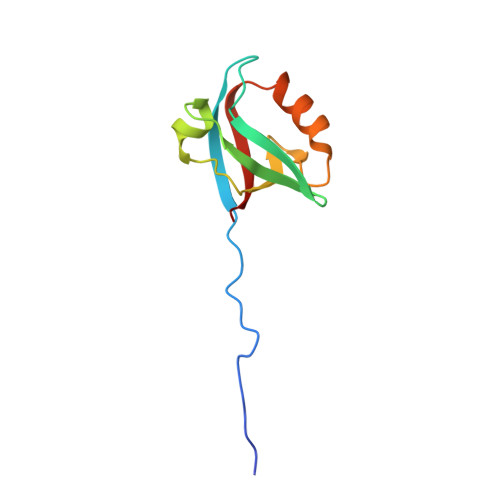
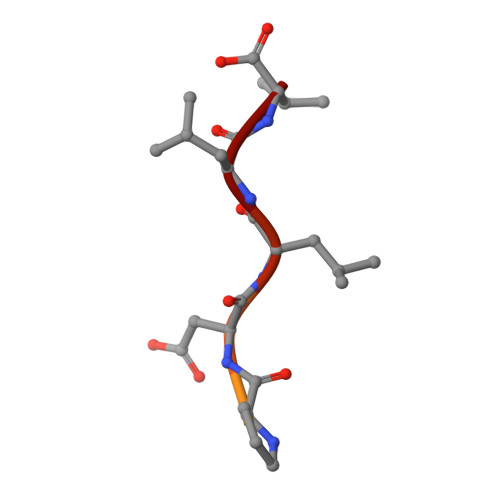
Interactions of Severe Acute Respiratory Syndrome Coronavirus 2 Protein E With Cell Junctions and Polarity PSD-95/Dlg/ZO-1-Containing Proteins.
Zhu, Y., Alvarez, F., Wolff, N., Mechaly, A., Brule, S., Neitthoffer, B., Etienne-Manneville, S., Haouz, A., Boeda, B., Caillet-Saguy, C.(2022) Front Microbiol 13: 829094-829094
- PubMed: 35283834
- DOI: https://doi.org/10.3389/fmicb.2022.829094
- Primary Citation of Related Structures:
7QCR, 7QCS, 7QCT - PubMed Abstract:
The C-terminus of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) protein E contains a PBM (PDZ-binding motif) targeting PDZ (PSD-95/Dlg/ZO-1) domains, which is identical to the PBM of SARS-CoV. The latter is involved in the pathogenicity of the virus. Recently, we identified 10 human PDZ-containing proteins showing significant interactions with SARS-CoV-2 protein E PBM. We selected several of them involved in cellular junctions and cell polarity (TJP1, PARD3, MLLT4, and LNX2) and MPP5/PALS1 previously shown to interact with SARS-CoV E PBM. Targeting cellular junctions and polarity components is a common strategy by viruses to hijack cell machinery to their advantage. In this study, we showed that these host PDZ domains TJP1, PARD3, MLLT4, LNX2, and MPP5/PALS1 interact in a PBM-dependent manner in vitro and colocalize with the full-length E protein in cellulo , sequestrating the PDZ domains to the Golgi compartment. We solved three crystal structures of complexes between human LNX2, MLLT4, and MPP5 PDZs and SARS-CoV-2 E PBM highlighting its binding preferences for several cellular targets. Finally, we showed different affinities for the PDZ domains with the original SARS-CoV-2 C-terminal sequence containing the PBM and the one of the beta variant that contains a mutation close to the PBM. The acquired mutations in the E protein localized near the PBM might have important effects both on the structure and the ion-channel activity of the E protein and on the host machinery targeted by the variants during the infection.
Organizational Affiliation:
Channel Receptors Unit, CNRS, UMR 3571, Institut Pasteur, Université de Paris, Paris, France.