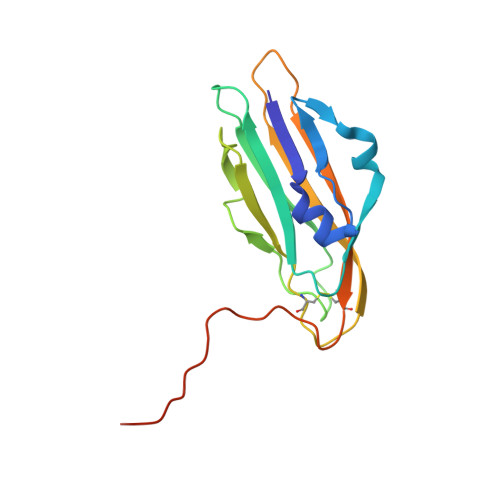
ROR and RYK extracellular region structures suggest that receptor tyrosine kinases have distinct WNT-recognition modes.
Shi, F., Mendrola, J.M., Sheetz, J.B., Wu, N., Sommer, A., Speer, K.F., Noordermeer, J.N., Kan, Z.Y., Perry, K., Englander, S.W., Stayrook, S.E., Fradkin, L.G., Lemmon, M.A.(2021) Cell Rep 37: 109834-109834
- PubMed: 34686333
- DOI: https://doi.org/10.1016/j.celrep.2021.109834
- Primary Citation of Related Structures:
7ME4, 7ME5 - PubMed Abstract:
WNTs play key roles in development and disease, signaling through Frizzled (FZD) seven-pass transmembrane receptors and numerous co-receptors including ROR and RYK family receptor tyrosine kinases (RTKs). We describe crystal structures and WNT-binding characteristics of extracellular regions from the Drosophila ROR and RYK orthologs Nrk (neurospecific receptor tyrosine kinase) and Derailed-2 (Drl-2), which bind WNTs though a FZD-related cysteine-rich domain (CRD) and WNT-inhibitory factor (WIF) domain respectively. Our crystal structures suggest that neither Nrk nor Drl-2 can accommodate the acyl chain typically attached to WNTs. The Nrk CRD contains a deeply buried bound fatty acid, unlikely to be exchangeable. The Drl-2 WIF domain lacks the lipid-binding site seen in WIF-1. We also find that recombinant DWnt-5 can bind Drosophila ROR and RYK orthologs despite lacking an acyl chain. Alongside analyses of WNT/receptor interaction sites, our structures provide further insight into how WNTs may recruit RTK co-receptors into signaling complexes.
Organizational Affiliation:
Department of Biochemistry and Biophysics, University of Pennsylvania Perelman School of Medicine, Philadelphia, PA 19104, USA; Graduate Group in Biochemistry and Molecular Biophysics, University of Pennsylvania Perelman School of Medicine, Philadelphia, PA 19104, USA.