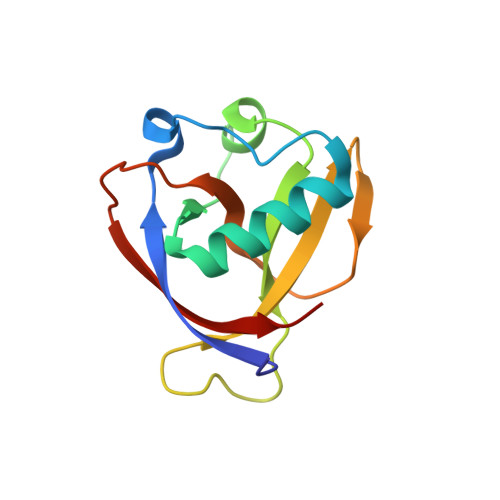
Structure of Nonstructural Protein 1 from SARS-CoV-2.
Clark, L.K., Green, T.J., Petit, C.M.(2021) J Virol 95
- PubMed: 33234675
- DOI: https://doi.org/10.1128/JVI.02019-20
- Primary Citation of Related Structures:
7K7P - PubMed Abstract:
The periodic emergence of novel coronaviruses (CoVs) represents an ongoing public health concern with significant health and financial burdens worldwide. The most recent occurrence originated in the city of Wuhan, China, where a novel coronavirus (severe acute respiratory syndrome coronavirus 2 [SARS-CoV-2]) emerged causing severe respiratory illness and pneumonia. The continual emergence of novel coronaviruses underscores the importance of developing effective vaccines as well as novel therapeutic options that target either viral functions or host factors recruited to support coronavirus replication. The CoV nonstructural protein 1 (nsp1) has been shown to promote cellular mRNA degradation, block host cell translation, and inhibit the innate immune response to virus infection. Interestingly, deletion of the nsp1-coding region in infectious clones prevented the virus from productively infecting cultured cells. Because of nsp1's importance in the CoV life cycle, it has been highlighted as a viable target for both antiviral therapy and vaccine development. However, the fundamental molecular and structural mechanisms that underlie nsp1 function remain poorly understood, despite its critical role in the viral life cycle. Here, we report the high-resolution crystal structure of the amino globular portion of SARS-CoV-2 nsp1 (residues 10 to 127) at 1.77-Å resolution. A comparison of our structure with the SARS-CoV-1 nsp1 structure reveals how mutations alter the conformation of flexible loops, inducing the formation of novel secondary structural elements and new surface features. Paired with the recently published structure of the carboxyl end of nsp1 (residues 148 to 180), our results provide the groundwork for future studies focusing on SARS-CoV-2 nsp1 structure and function during the viral life cycle. IMPORTANCE Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is the causative agent of the COVID-19 pandemic. One protein known to play a critical role in the coronavirus life cycle is nonstructural protein 1 (nsp1). As such, it has been highlighted in numerous studies as a target for both the development of antivirals and the design of live-attenuated vaccines. Here, we report the high-resolution crystal structure of nsp1 derived from SARS-CoV-2 at 1.77-Å resolution. This structure will facilitate future studies focusing on understanding the relationship between structure and function for nsp1. In turn, understanding these structure-function relationships will allow nsp1 to be fully exploited as a target for both antiviral development and vaccine design.
Organizational Affiliation:
Department of Biochemistry and Molecular Genetics, University of Alabama at Birmingham School of Medicine, Birmingham, Alabama, USA.