Crystal structure of SARS-CoV-2 Y453F-RBD bound to mink ACE2
Wang, X.Q., Ding, Q., Lan, J., Ren, W.L.To be published.
Experimental Data Snapshot
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| mink ACE2 | A, C [auth B], E [auth D] | 597 | Neogale vison | Mutation(s): 0 | 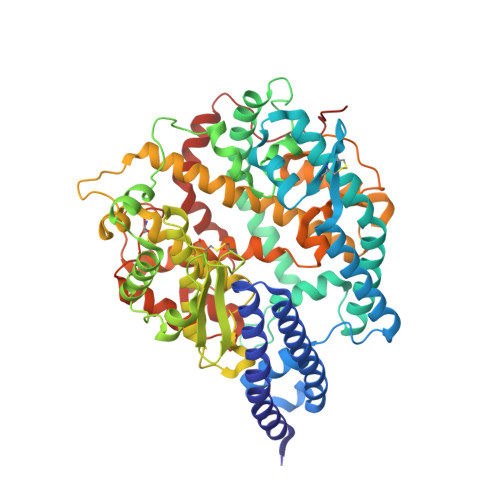 |
UniProt | |||||
Find proteins for A0A8C7BTF2 (Neovison vison) Explore A0A8C7BTF2 Go to UniProtKB: A0A8C7BTF2 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A0A8C7BTF2 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Spike protein S1 | B [auth E], D [auth C], F | 197 | Severe acute respiratory syndrome coronavirus 2 | Mutation(s): 1 Gene Names: S, 2 | 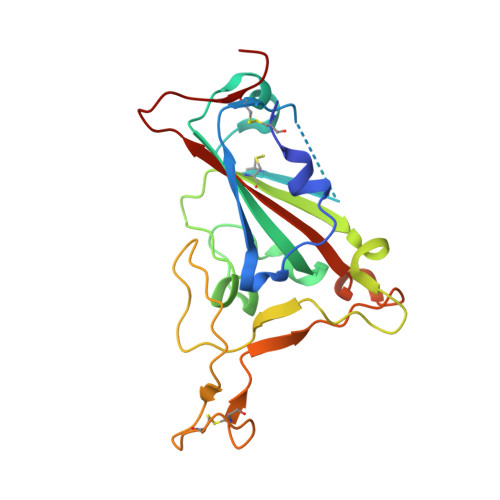 |
UniProt | |||||
Find proteins for P0DTC2 (Severe acute respiratory syndrome coronavirus 2) Explore P0DTC2 Go to UniProtKB: P0DTC2 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0DTC2 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NAG Query on NAG | J [auth A] K [auth E] L [auth B] M [auth B] N [auth B] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 290.185 | α = 90 |
| b = 130.185 | β = 90 |
| c = 136.887 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| PDB_EXTRACT | data extraction |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |
| MOLREP | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Not funded | -- |