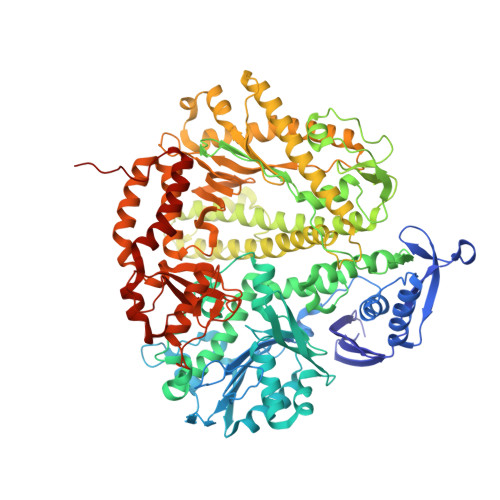
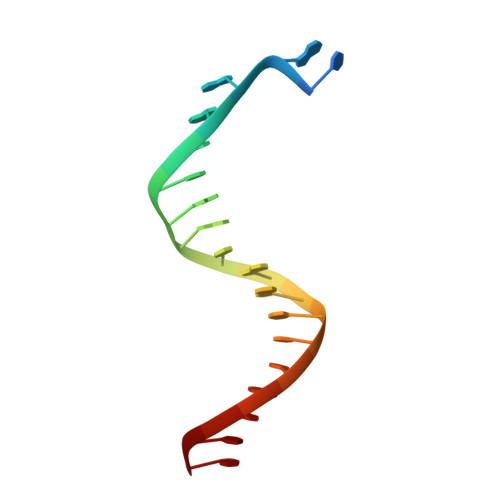
Structure of New Binary and Ternary DNA Polymerase Complexes From Bacteriophage RB69.
Park, J., Youn, H.S., An, J.Y., Lee, Y., Eom, S.H., Wang, J.(2021) Front Mol Biosci 8: 704813-704813
- PubMed: 34869578
- DOI: https://doi.org/10.3389/fmolb.2021.704813
- Primary Citation of Related Structures:
7F4Y - PubMed Abstract:
DNA polymerase plays a critical role in passing the genetic information of any living organism to its offspring. DNA polymerase from enterobacteria phage RB69 (RB69pol) has both polymerization and exonuclease activities and has been extensively studied as a model system for B-family DNA polymerases. Many binary and ternary complex structures of RB69pol are known, and they all contain a single polymerase-primer/template (P/T) DNA complex. Here, we report a crystal structure of the exonuclease-deficient RB69pol with the P/T duplex in a dimeric form at a resolution of 2.2 Å. The structure includes one new closed ternary complex with a single divalent metal ion bound and one new open binary complex in the pre-insertion state with a vacant dNTP-binding pocket. These complexes suggest that initial binding of the correct dNTP in the open state is much weaker than expected and that initial binding of the second divalent metal ion in the closed state is also much weaker than measured. Additional conformational changes are required to convert these complexes to high-affinity states. Thus, the measured affinities for the correct incoming dNTP and divalent metal ions are average values from many conformationally distinctive states. Our structure provides new insights into the order of the complex assembly involving two divalent metal ions. The biological relevance of specific interactions observed between one RB69pol and the P/T duplex bound to the second RB69pol observed within this dimeric complex is discussed.
Organizational Affiliation:
School of Life Sciences, Gwangju Institute of Science and Technology (GIST), Gwangju, South Korea.