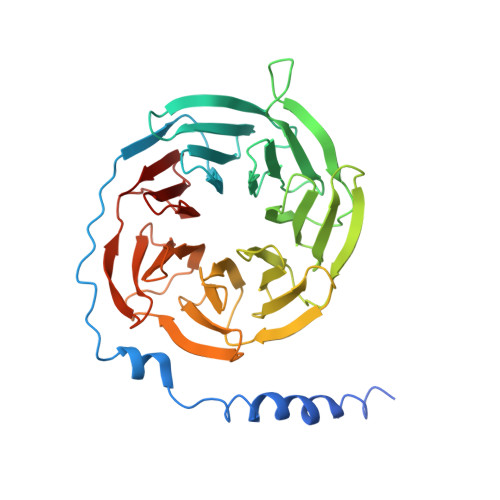
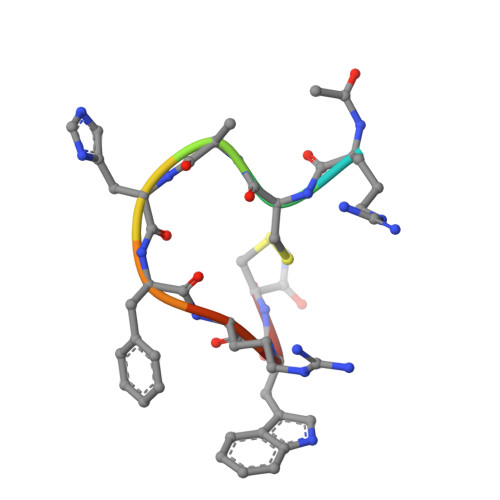
Structure reveals the activation mechanism of the MC4 receptor to initiate satiation signaling.
Israeli, H., Degtjarik, O., Fierro, F., Chunilal, V., Gill, A.K., Roth, N.J., Botta, J., Prabahar, V., Peleg, Y., Chan, L.F., Ben-Zvi, D., McCormick, P.J., Niv, M.Y., Shalev-Benami, M.(2021) Science 372: 808-814
- PubMed: 33858992
- DOI: https://doi.org/10.1126/science.abf7958
- Primary Citation of Related Structures:
7AUE - PubMed Abstract:
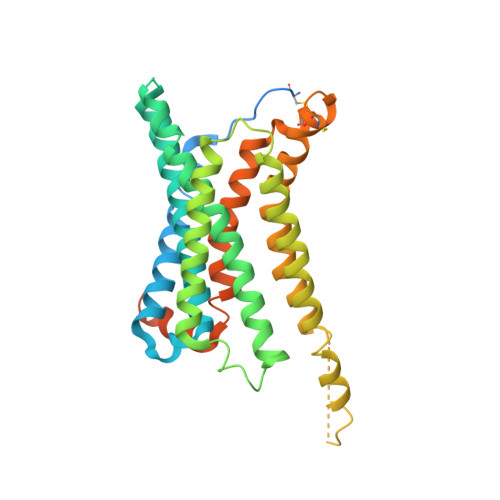
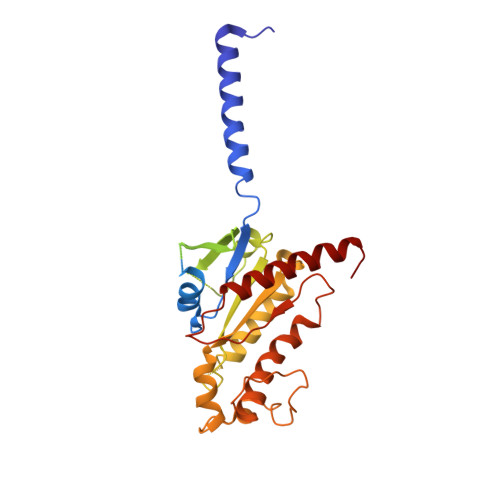
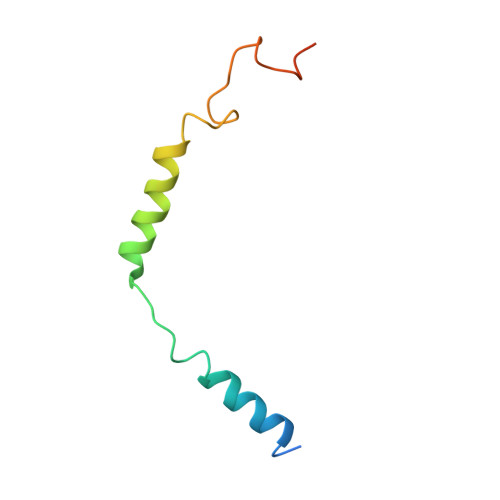
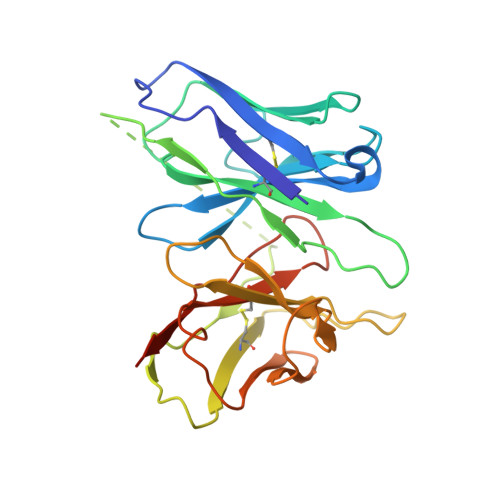
Obesity is a global epidemic that causes morbidity and impaired quality of life. The melanocortin receptor 4 (MC4R) is at the crux of appetite, energy homeostasis, and body-weight control in the central nervous system and is a prime target for anti-obesity drugs. Here, we present the cryo-electron microscopy (cryo-EM) structure of the human MC4R-G s signaling complex bound to the agonist setmelanotide, a cyclic peptide recently approved for the treatment of obesity. The work reveals the mechanism of MC4R activation, highlighting a molecular switch that initiates satiation signaling. In addition, our findings indicate that calcium (Ca 2+ ) is required for agonist, but not antagonist, efficacy. These results fill a gap in the understanding of MC4R activation and could guide the design of future weight-management drugs.
Organizational Affiliation:
Department of Chemical and Structural Biology, Weizmann Institute of Science, Rehovot 7610001, Israel.