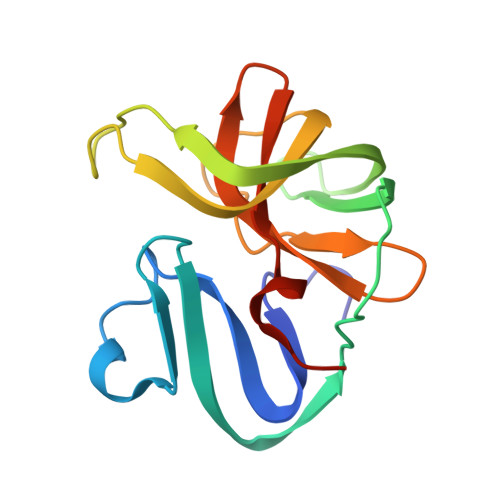
Defining substrate selection by rhinoviral 2A proteinase through its crystal structure with the inhibitor zVAM.fmk.
Deutschmann-Olek, K.M., Yue, W.W., Bezerra, G.A., Skern, T.(2021) Virology 562: 128-141
- PubMed: 34315103
- DOI: https://doi.org/10.1016/j.virol.2021.07.008
- Primary Citation of Related Structures:
7ARA - PubMed Abstract:
Picornavirus family members cause disease in humans. Human rhinoviruses (RV), the main causative agents of the common cold, increase the severity of asthma and COPD; hence, effective agents against RVs are required. The 2A proteinase (2A pro ), found in all enteroviruses, represents an attractive target; inactivating mutations in poliovirus 2A pro result in an extension of the VP1 protein preventing infectious virion assembly. Variations in sequence and substrate specificity on eIF4G isoforms between RV 2A pro of genetic groups A and B hinder 2A pro as drug targets. Here, we demonstrate that although RV-A2 and RV-B4 2A pro cleave the substrate GAB1 at different sites, the 2A pro from both groups cleave equally efficiently an artificial site containing P1 methionine. We determined the RV-A2 2A pro structure complexed with zVAM.fmk, containing P1 methionine. Analysis of this first 2A pro -inhibitor complex reveals a conserved hydrophobic P4 pocket among enteroviral 2A pro as a potential target for broad-spectrum anti-enteroviral inhibitors.
Organizational Affiliation:
Department of Medical Biochemistry, Max Perutz Labs, Vienna Biocenter, Medical University of Vienna, A-1030, Vienna, Austria.