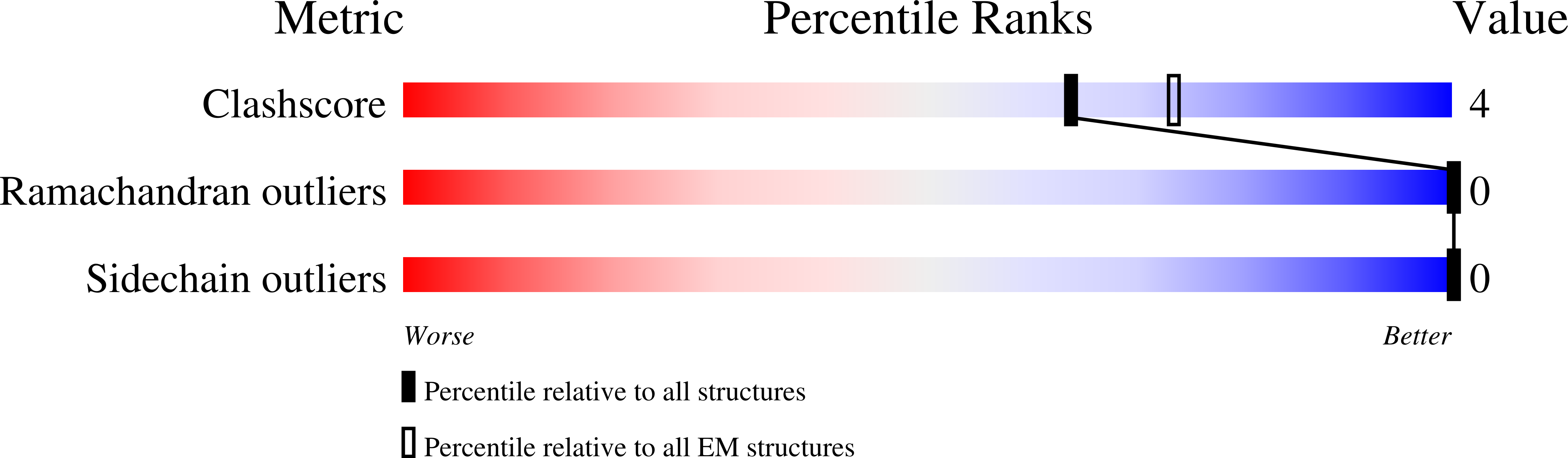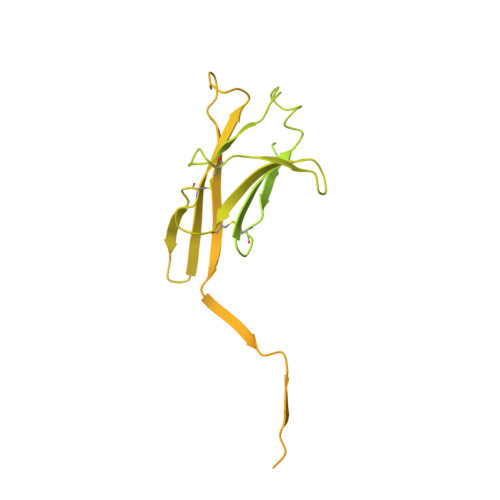The cryo-EM structure of the human uromodulin filament core reveals a unique assembly mechanism.
Stanisich, J.J., Zyla, D.S., Afanasyev, P., Xu, J., Kipp, A., Olinger, E., Devuyst, O., Pilhofer, M., Boehringer, D., Glockshuber, R.(2020) Elife 9
- PubMed: 32815518
- DOI: https://doi.org/10.7554/eLife.60265
- Primary Citation of Related Structures:
6ZS5, 6ZYA - PubMed Abstract:
The glycoprotein uromodulin (UMOD) is the most abundant protein in human urine and forms filamentous homopolymers that encapsulate and aggregate uropathogens, promoting pathogen clearance by urine excretion. Despite its critical role in the innate immune response against urinary tract infections, the structural basis and mechanism of UMOD polymerization remained unknown. Here, we present the cryo-EM structure of the UMOD filament core at 3.5 Å resolution, comprised of the bipartite zona pellucida (ZP) module in a helical arrangement with a rise of ~65 Å and a twist of ~180°. The immunoglobulin-like ZPN and ZPC subdomains of each monomer are separated by a long linker that interacts with the preceding ZPC and following ZPN subdomains by β-sheet complementation. The unique filament architecture suggests an assembly mechanism in which subunit incorporation could be synchronized with proteolytic cleavage of the C-terminal pro-peptide that anchors assembly-incompetent UMOD precursors to the membrane.
Organizational Affiliation:
Institute of Molecular Biology & Biophysics, ETH Zurich, Zurich, Switzerland.