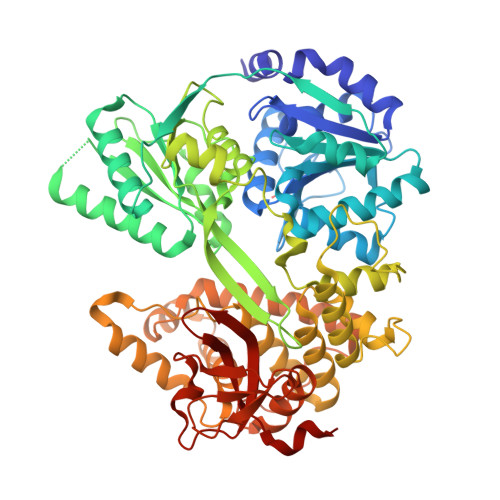
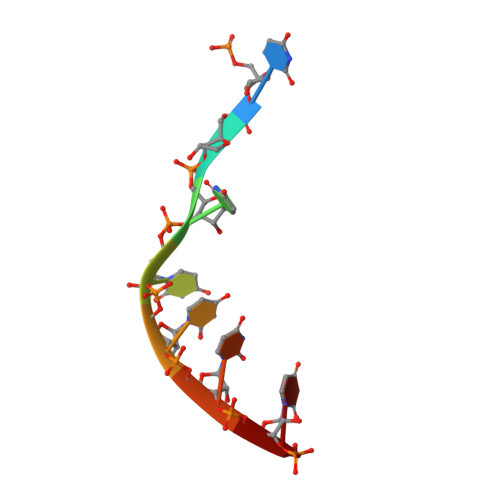
The structure of Prp2 bound to RNA and ADP-BeF 3 - reveals structural features important for RNA unwinding by DEAH-box ATPases.
Hamann, F., Zimmerningkat, L.C., Becker, R.A., Garbers, T.B., Neumann, P., Hub, J.S., Ficner, R.(2021) Acta Crystallogr D Struct Biol 77: 496-509
- PubMed: 33825710
- DOI: https://doi.org/10.1107/S2059798321001194
- Primary Citation of Related Structures:
6ZM2 - PubMed Abstract:
Noncoding intron sequences present in precursor mRNAs need to be removed prior to translation, and they are excised via the spliceosome, a multimegadalton molecular machine composed of numerous protein and RNA components. The DEAH-box ATPase Prp2 plays a crucial role during pre-mRNA splicing as it ensures the catalytic activation of the spliceosome. Despite high structural similarity to other spliceosomal DEAH-box helicases, Prp2 does not seem to function as an RNA helicase, but rather as an RNA-dependent ribonucleoprotein particle-modifying ATPase. Recent crystal structures of the spliceosomal DEAH-box ATPases Prp43 and Prp22, as well as of the related RNA helicase MLE, in complex with RNA have contributed to a better understanding of how RNA binding and processivity might be achieved in this helicase family. In order to shed light onto the divergent manner of function of Prp2, an N-terminally truncated construct of Chaetomium thermophilum Prp2 was crystallized in the presence of ADP-BeF 3 - and a poly-U 12 RNA. The refined structure revealed a virtually identical conformation of the helicase core compared with the ADP-BeF 3 - - and RNA-bound structure of Prp43, and only a minor shift of the C-terminal domains. However, Prp2 and Prp43 differ in the hook-loop and a loop of the helix-bundle domain, which interacts with the hook-loop and evokes a different RNA conformation immediately after the 3' stack. On replacing these loop residues in Prp43 by the Prp2 sequence, the unwinding activity of Prp43 was abolished. Furthermore, a putative exit tunnel for the γ-phosphate after ATP hydrolysis could be identified in one of the Prp2 structures.
Organizational Affiliation:
Department of Molecular Structural Biology, Institute of Microbiology and Genetics, Göttingen Center for Molecular Biosciences (GZMB), Georg-August-University Göttingen, Justus-von-Liebig-Weg 11, 37077 Göttingen, Germany.