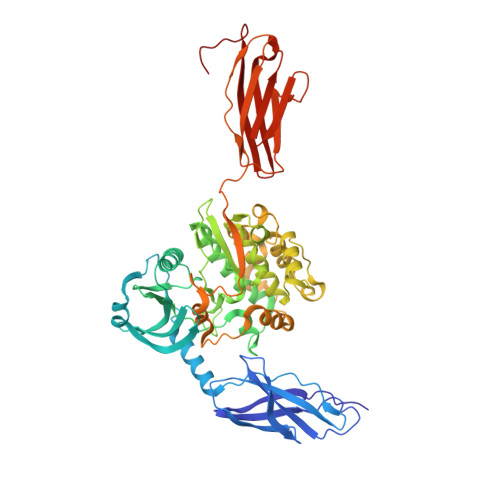
Titin kinase ubiquitination aligns autophagy receptors with mechanical signals in the sarcomere.
Bogomolovas, J., Fleming, J.R., Franke, B., Manso, B., Simon, B., Gasch, A., Markovic, M., Brunner, T., Knoll, R., Chen, J., Labeit, S., Scheffner, M., Peter, C., Mayans, O.(2021) EMBO Rep 22: e48018-e48018
- PubMed: 34402565
- DOI: https://doi.org/10.15252/embr.201948018
- Primary Citation of Related Structures:
6YGN - PubMed Abstract:
Striated muscle undergoes remodelling in response to mechanical and physiological stress, but little is known about the integration of such varied signals in the myofibril. The interaction of the elastic kinase region from sarcomeric titin (A168-M1) with the autophagy receptors Nbr1/p62 and MuRF E3 ubiquitin ligases is well suited to link mechanosensing with the trophic response of the myofibril. To investigate the mechanisms of signal cross-talk at this titin node, we elucidated its 3D structure, analysed its response to stretch using steered molecular dynamics simulations and explored its functional relation to MuRF1 and Nbr1/p62 using cellular assays. We found that MuRF1-mediated ubiquitination of titin kinase promotes its scaffolding of Nbr1/p62 and that the process can be dynamically down-regulated by the mechanical unfolding of a linker sequence joining titin kinase with the MuRF1 receptor site in titin. We propose that titin ubiquitination is sensitive to the mechanical state of the sarcomere, the regulation of sarcomere targeting by Nbr1/p62 being a functional outcome. We conclude that MuRF1/Titin Kinase/Nbr1/p62 constitutes a distinct assembly that predictably promotes sarcomere breakdown in inactive muscle.
Organizational Affiliation:
Department of Medicine, School of Medicine, University of California, San Diego, La Jolla, CA, USA.