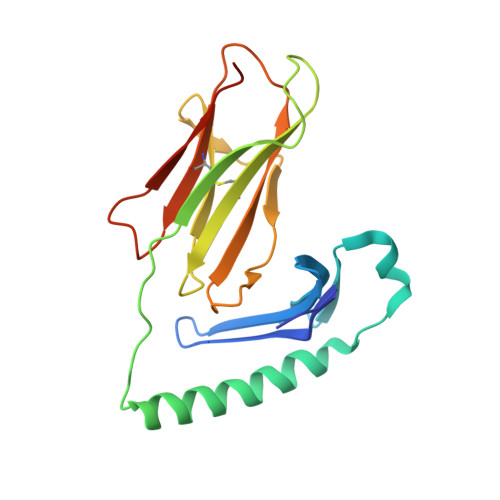
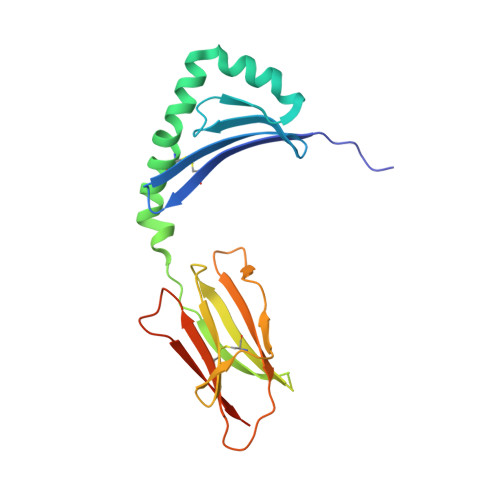
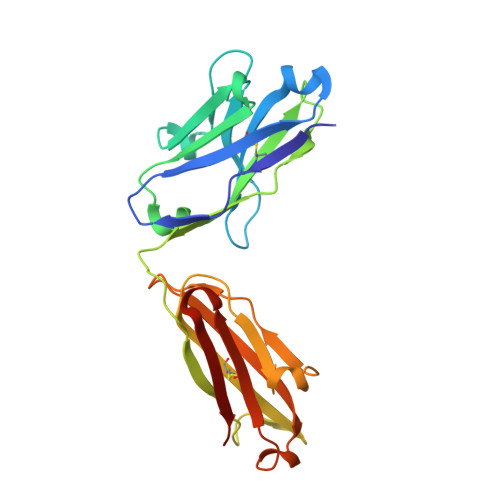
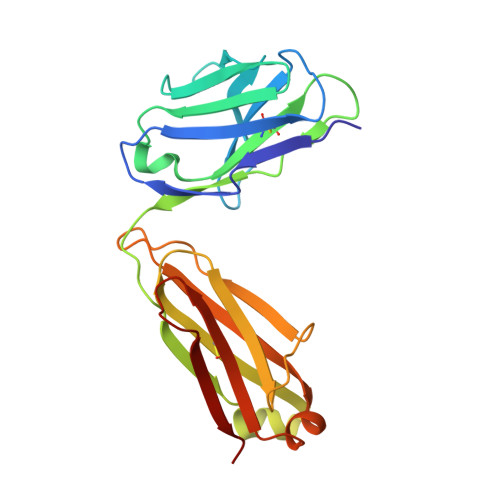
A high-affinity human TCR-like antibody detects celiac disease gluten peptide-MHC complexes and inhibits T cell activation.
Frick, R., Hoydahl, L.S., Petersen, J., du Pre, M.F., Kumari, S., Berntsen, G., Dewan, A.E., Jeliazkov, J.R., Gunnarsen, K.S., Frigstad, T., Vik, E.S., Llerena, C., Lundin, K.E.A., Yaqub, S., Jahnsen, J., Gray, J.J., Rossjohn, J., Sollid, L.M., Sandlie, I., Loset, G.A.(2021) Sci Immunol 6
- PubMed: 34417258
- DOI: https://doi.org/10.1126/sciimmunol.abg4925
- Primary Citation of Related Structures:
6XP6 - PubMed Abstract:
Antibodies specific for peptides bound to human leukocyte antigen (HLA) molecules are valuable tools for studies of antigen presentation and may have therapeutic potential. Here, we generated human T cell receptor (TCR)-like antibodies toward the immunodominant signature gluten epitope DQ2.5-glia-α2 in celiac disease (CeD). Phage display selection combined with secondary targeted engineering was used to obtain highly specific antibodies with picomolar affinity. The crystal structure of a Fab fragment of the lead antibody 3.C11 in complex with HLA-DQ2.5:DQ2.5-glia-α2 revealed a binding geometry and interaction mode highly similar to prototypic TCRs specific for the same complex. Assessment of CeD biopsy material confirmed disease specificity and reinforced the notion that abundant plasma cells present antigen in the inflamed CeD gut. Furthermore, 3.C11 specifically inhibited activation and proliferation of gluten-specific CD4 + T cells in vitro and in HLA-DQ2.5 humanized mice, suggesting a potential for targeted intervention without compromising systemic immunity.
Organizational Affiliation:
Centre for Immune Regulation and Department of Immunology, University of Oslo and Oslo University Hospital-Rikshospitalet, Oslo, Norway.