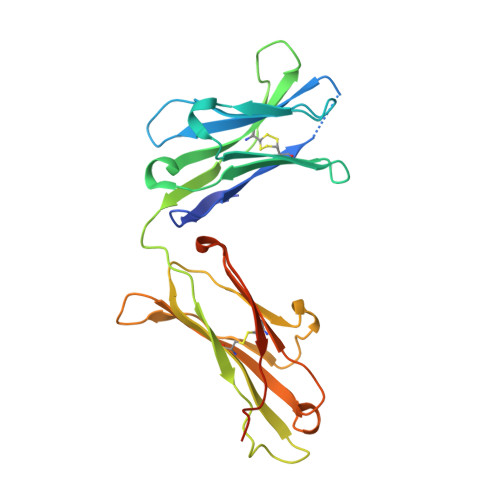
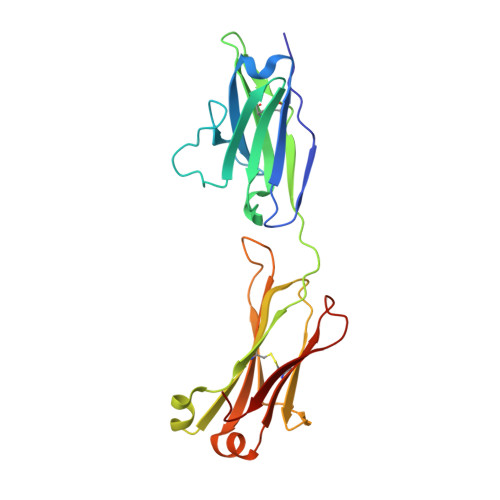
Crystal Structure of a Bivalent Antibody Fab Fragment.
Shahid, S., Gao, M., Travis Gallagher, D., Pozharski, E., Brinson, R.G., Keck, Z.Y., Foung, S.K.H., Fuerst, T.R., Mariuzza, R.A.(2020) J Mol Biol 433: 166714-166714
- PubMed: 33220264
- DOI: https://doi.org/10.1016/j.jmb.2020.11.013
- Primary Citation of Related Structures:
6X9X - PubMed Abstract:
We determined the crystal structure to 1.8 Å resolution of the Fab fragment of an affinity-matured human monoclonal antibody (HC84.26.5D) that recognizes the E2 envelope glycoprotein of hepatitis C virus (HCV). Unlike conventional Fabs, which are monovalent monomers, Fab HC84.26.5D assembles into a bivalent domain-swapped dimer in which the two V L /V H modules are separated by ~25 Å. In solution, Fab HC84.26.5D exists predominantly as a dimer (~80%) in equilibrium with the monomeric form of the Fab (~20%). Dimerization is mediated entirely by deletion of a single residue, V H Ser113 (Kabat numbering), in the elbow region linking the V H and C H 1 domains. In agreement with the crystal structure, dimeric Fab HC84.26.5D is able to bind two HCV E2 molecules in solution. This is only the second example of a domain-swapped Fab dimer from among >3000 Fab crystal structures determined to date. Moreover, the architecture of the doughnut-shaped Fab HC84.26.5D dimer is completely different from that of the previously reported Fab 2G12 dimer. We demonstrate that the highly identifiable shape of dimeric Fab HC84.26.5D makes it useful as a fiducial marker for single-particle cryoEM analysis of HCV E2. Bivalent domain-swapped Fab dimers engineered on the basis of HC84.26.5D may also serve as a means of doubling the effective size of conventional Fab-protein complexes for cryoEM.
Organizational Affiliation:
University of Maryland Institute for Bioscience and Biotechnology Research, Rockville, MD 20850, USA; Department of Cell Biology and Molecular Genetics, University of Maryland, College Park, MD 20742, USA.