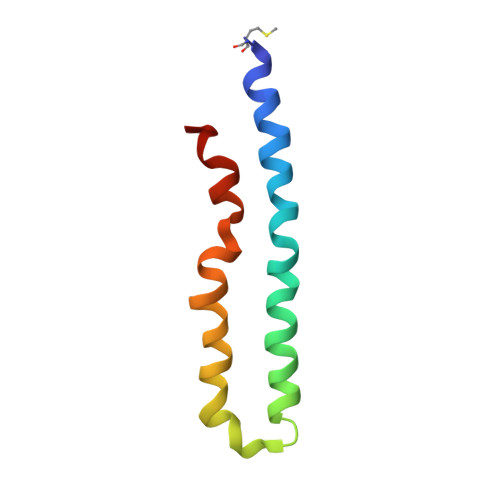
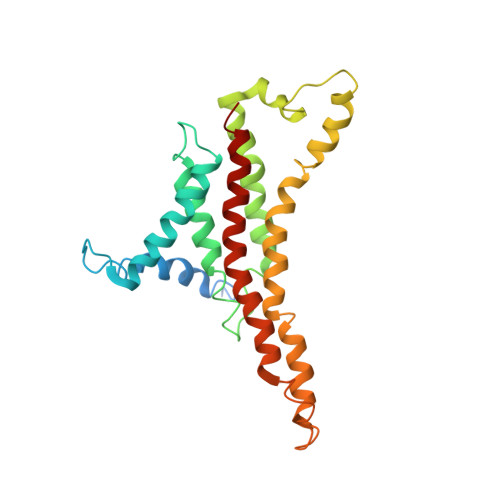
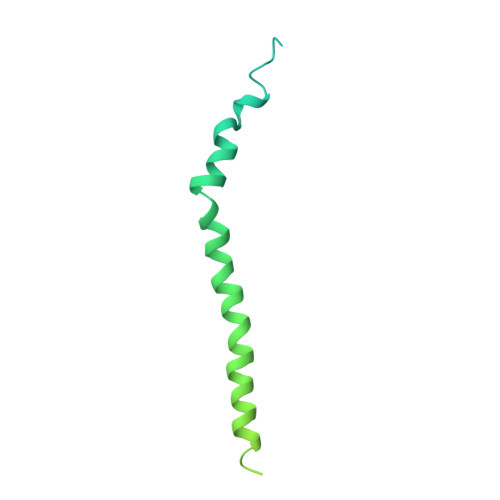
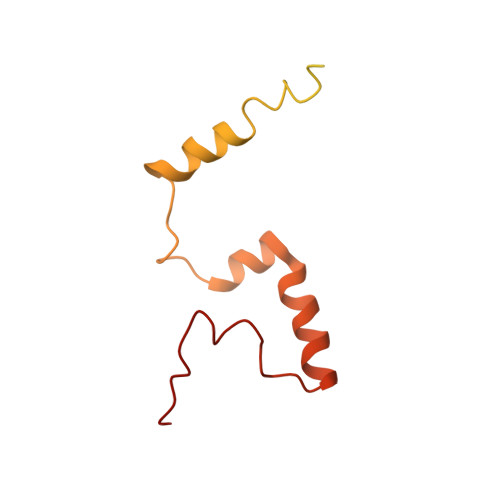
Bedaquiline inhibits the yeast and human mitochondrial ATP synthases.
Luo, M., Zhou, W., Patel, H., Srivastava, A.P., Symersky, J., Bonar, M.M., Faraldo-Gomez, J.D., Liao, M., Mueller, D.M.(2020) Commun Biol 3: 452-452
- PubMed: 32814813
- DOI: https://doi.org/10.1038/s42003-020-01173-z
- Primary Citation of Related Structures:
6WTD - PubMed Abstract:
Bedaquiline (BDQ, Sirturo) has been approved to treat multidrug resistant forms of Mycobacterium tuberculosis. Prior studies suggested that BDQ was a selective inhibitor of the ATP synthase from M. tuberculosis. However, Sirturo treatment leads to an increased risk of cardiac arrhythmias and death, raising the concern that this adverse effect results from inhibition at a secondary site. Here we show that BDQ is a potent inhibitor of the yeast and human mitochondrial ATP synthases. Single-particle cryo-EM reveals that the site of BDQ inhibition partially overlaps with that of the inhibitor oligomycin. Molecular dynamics simulations indicate that the binding mode of BDQ to this site is similar to that previously seen for a mycobacterial enzyme, explaining the observed lack of selectivity. We propose that derivatives of BDQ ought to be made to increase its specificity toward the mycobacterial enzyme and thereby reduce the side effects for patients that are treated with Sirturo.
Organizational Affiliation:
Department of Cell Biology, Harvard Medical School, 250 Longwood Avenue, Boston, MA, 02115, USA.