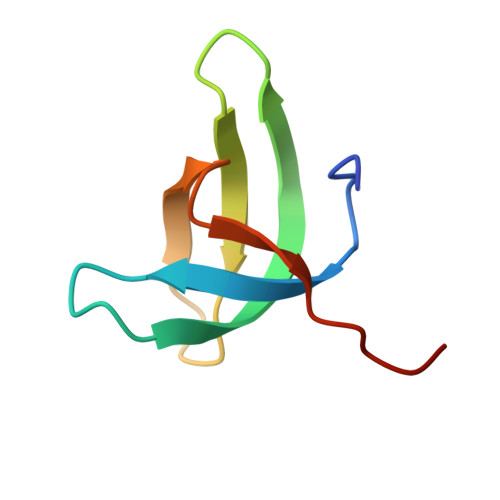
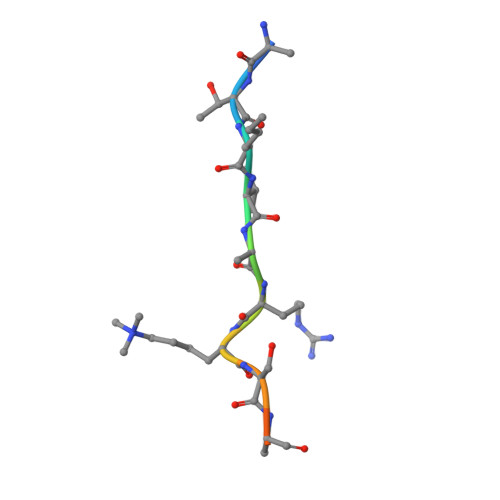
Structural basis for histone variant H3tK27me3 recognition by PHF1 and PHF19.
Dong, C., Nakagawa, R., Oyama, K., Yamamoto, Y., Zhang, W., Dong, A., Li, Y., Yoshimura, Y., Kamiya, H., Nakayama, J.I., Ueda, J., Min, J.(2020) Elife 9
- PubMed: 32869745
- DOI: https://doi.org/10.7554/eLife.58675
- Primary Citation of Related Structures:
6WAT, 6WAU, 6WAV - PubMed Abstract:
The Polycomb repressive complex 2 (PRC2) is a multicomponent histone H3K27 methyltransferase complex, best known for silencing the Hox genes during embryonic development. The Polycomb-like proteins PHF1, MTF2, and PHF19 are critical components of PRC2 by stimulating its catalytic activity in embryonic stem cells. The Tudor domains of PHF1/19 have been previously shown to be readers of H3K36me3 in vitro. However, some other studies suggest that PHF1 and PHF19 co-localize with the H3K27me3 mark but not H3K36me3 in cells. Here, we provide further evidence that PHF1 co-localizes with H3t in testis and its Tudor domain preferentially binds to H3tK27me3 over canonical H3K27me3 in vitro. Our complex structures of the Tudor domains of PHF1 and PHF19 with H3tK27me3 shed light on the molecular basis for preferential recognition of H3tK27me3 by PHF1 and PHF19 over canonical H3K27me3, implicating that H3tK27me3 might be a physiological ligand of PHF1/19.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, School of Basic Medical Sciences, Tianjin Medical University, Tianjin, China.