DNA Polymerase Mu, 8-oxorGTP:At Reaction State Ternary Complex, 50 mM Mn2+ (30 min)
Jamsen, J.A., Sassa, A., Shock, D.D., Beard, W.A., Wilson, S.H.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Currently 6VF4 does not have a validation slider image.
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| DNA-directed DNA/RNA polymerase mu | 356 | Homo sapiens | Mutation(s): 0 Gene Names: POLM, polmu EC: 2.7.7.7 | 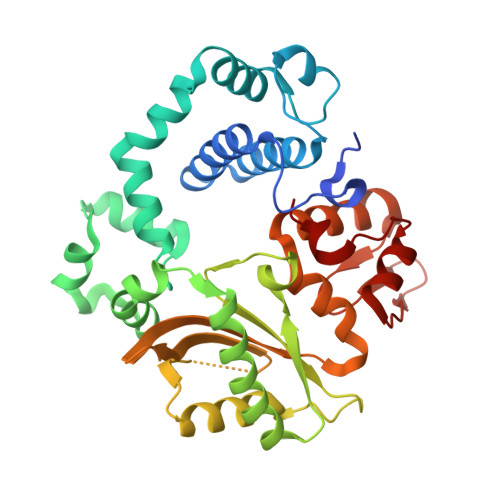 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for Q9NP87 (Homo sapiens) Explore Q9NP87 Go to UniProtKB: Q9NP87 | |||||
PHAROS: Q9NP87 GTEx: ENSG00000122678 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9NP87 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Find similar nucleic acids by: Sequence | 3D Structure
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | Organism | Image | |
| DNA (5'-D(*CP*GP*GP*CP*AP*TP*AP*CP*G)-3') | B [auth T] | 9 | synthetic construct | 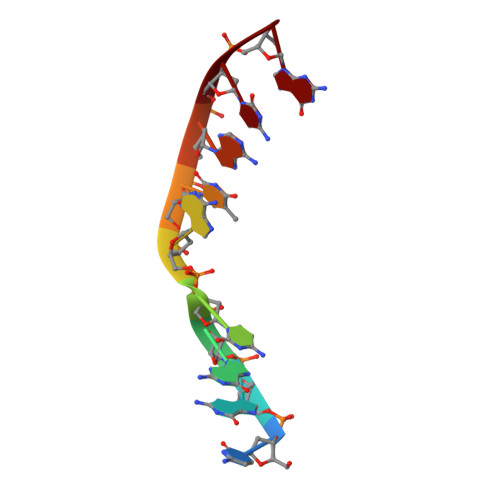 | |
Sequence AnnotationsExpand | |||||
| |||||
Find similar nucleic acids by: Sequence | 3D Structure
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | Organism | Image | |
| DNA (5'-D(*CP*GP*TP*AP*(8GM))-3') | C [auth P] | 5 | synthetic construct | 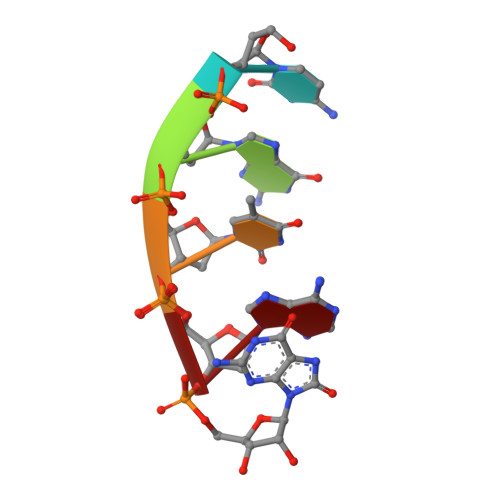 | |
Sequence AnnotationsExpand | |||||
| |||||
Find similar nucleic acids by: Sequence | 3D Structure
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | Organism | Image | |
| DNA (5'-D(P*GP*CP*CP*G)-3') | 4 | synthetic construct | 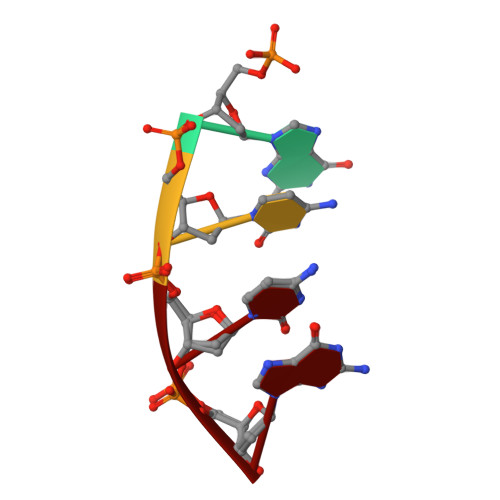 | ||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 7 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| 8GT (Subject of Investigation/LOI) Query on 8GT | O [auth A] | 8-OXO-GUANOSINE-5'-TRIPHOSPHATE C10 H16 N5 O15 P3 JCHLKIQZUXYLPW-UMMCILCDSA-N |  | ||
| EPE Query on EPE | L [auth A] | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID C8 H18 N2 O4 S JKMHFZQWWAIEOD-UHFFFAOYSA-N |  | ||
| PPV (Subject of Investigation/LOI) Query on PPV | J [auth A] | PYROPHOSPHATE H4 O7 P2 XPPKVPWEQAFLFU-UHFFFAOYSA-N |  | ||
| DTT Query on DTT | K [auth A] | 2,3-DIHYDROXY-1,4-DITHIOBUTANE C4 H10 O2 S2 VHJLVAABSRFDPM-IMJSIDKUSA-N |  | ||
| EDO Query on EDO | M [auth A], N [auth A], Q [auth P] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| MN (Subject of Investigation/LOI) Query on MN | E [auth A], F [auth A], G [auth A], H [auth A], P [auth T] | MANGANESE (II) ION Mn WAEMQWOKJMHJLA-UHFFFAOYSA-N |  | ||
| NA Query on NA | I [auth A] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 60.085 | α = 90 |
| b = 68.757 | β = 90 |
| c = 110.531 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |
| PDB_EXTRACT | data extraction |
| PHENIX | phasing |
Currently 6VF4 does not have a validation slider image.
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute of Environmental Health Sciences (NIH/NIEHS) | United States | Z01-ES050158 |
| National Institutes of Health/National Institute of Environmental Health Sciences (NIH/NIEHS) | United States | Z01-10ES050161 |
| National Institutes of Health/National Institute of Environmental Health Sciences (NIH/NIEHS) | United States | 1K99ES029572-01 |
| Japan Society for the Promotion of Science (JSPS) | Japan | 16K16195 |