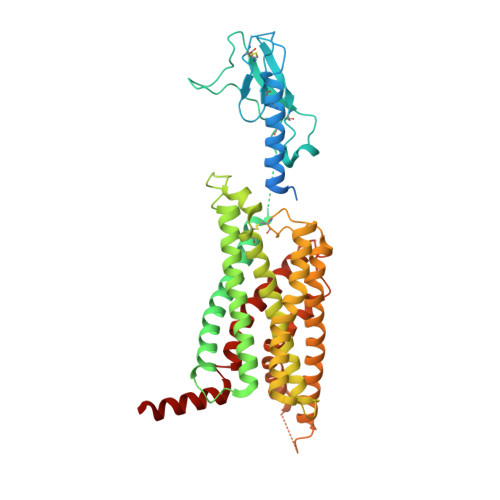
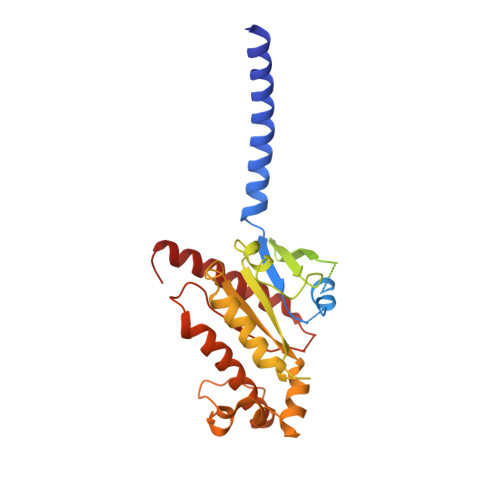
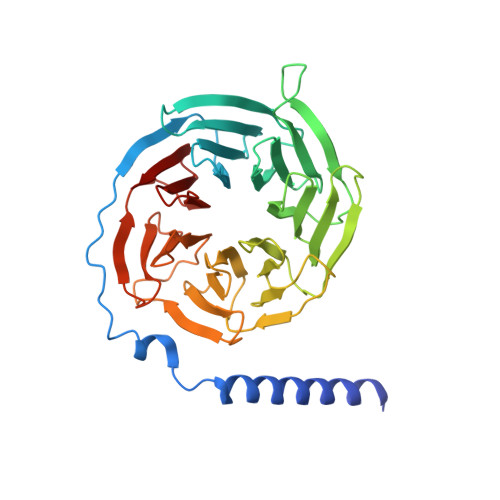
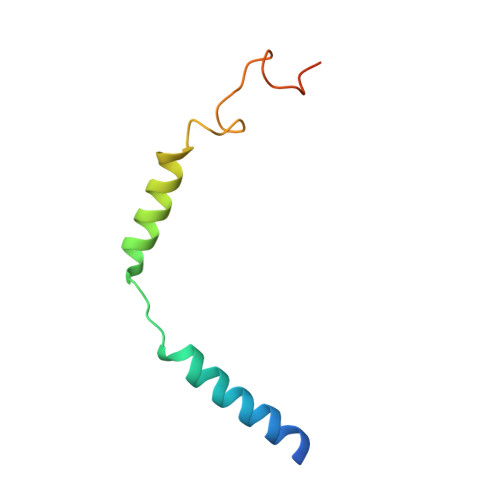
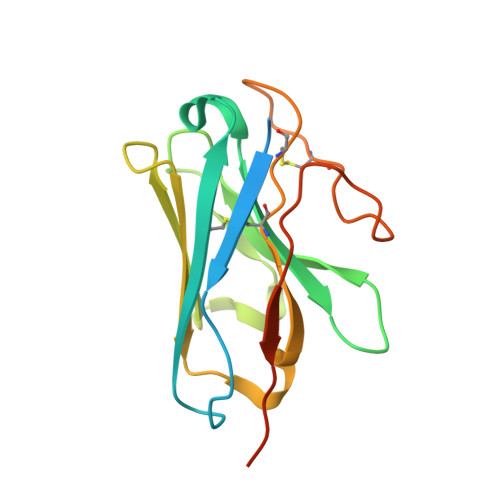
Structural insights into probe-dependent positive allosterism of the GLP-1 receptor.
Bueno, A.B., Sun, B., Willard, F.S., Feng, D., Ho, J.D., Wainscott, D.B., Showalter, A.D., Vieth, M., Chen, Q., Stutsman, C., Chau, B., Ficorilli, J., Agejas, F.J., Cumming, G.R., Jimenez, A., Rojo, I., Kobilka, T.S., Kobilka, B.K., Sloop, K.W.(2020) Nat Chem Biol 16: 1105-1110
- PubMed: 32690941
- DOI: https://doi.org/10.1038/s41589-020-0589-7
- Primary Citation of Related Structures:
6VCB - PubMed Abstract:
Drugs that promote the association of protein complexes are an emerging therapeutic strategy. We report discovery of a G protein-coupled receptor (GPCR) ligand that stabilizes an active state conformation by cooperatively binding both the receptor and orthosteric ligand, thereby acting as a 'molecular glue'. LSN3160440 is a positive allosteric modulator of the GLP-1R optimized to increase the affinity and efficacy of GLP-1(9-36), a proteolytic product of GLP-1(7-36). The compound enhances insulin secretion in a glucose-, ligand- and GLP-1R-dependent manner. Cryo-electron microscopy determined the structure of the GLP-1R bound to LSN3160440 in complex with GLP-1 and heterotrimeric G s . The modulator binds high in the helical bundle at an interface between TM1 and TM2, allowing access to the peptide ligand. Pharmacological characterization showed strong probe dependence of LSN3160440 for GLP-1(9-36) versus oxyntomodulin that is driven by a single residue. Our findings expand protein-protein modulation drug discovery to uncompetitive, active state stabilizers for peptide hormone receptors.
Organizational Affiliation:
Lilly, S.A., Alcobendas, Spain.