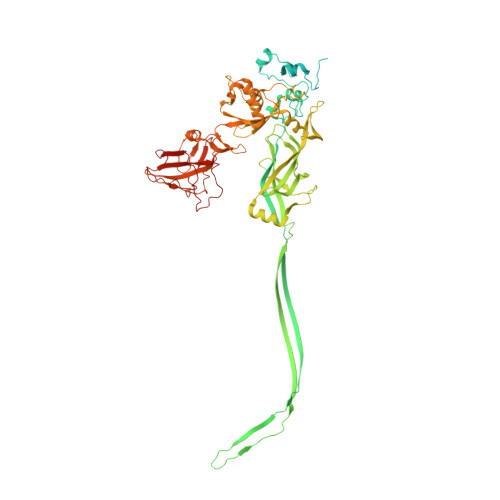
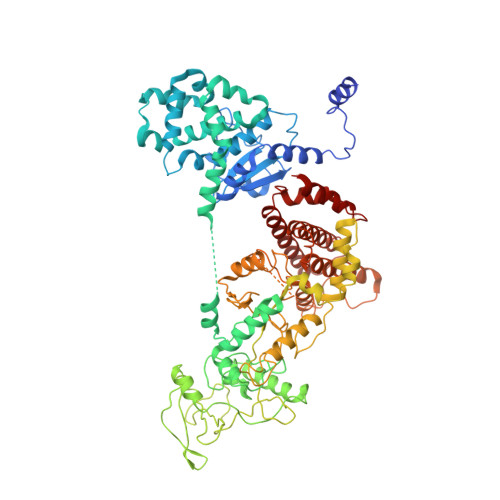
Atomic structures of anthrax toxin protective antigen channels bound to partially unfolded lethal and edema factors.
Hardenbrook, N.J., Liu, S., Zhou, K., Ghosal, K., Hong Zhou, Z., Krantz, B.A.(2020) Nat Commun 11: 840-840
- PubMed: 32047164
- DOI: https://doi.org/10.1038/s41467-020-14658-6
- Primary Citation of Related Structures:
6PSN, 6UZB, 6UZD, 6UZE - PubMed Abstract:
Following assembly, the anthrax protective antigen (PA) forms an oligomeric translocon that unfolds and translocates either its lethal factor (LF) or edema factor (EF) into the host cell. Here, we report the cryo-EM structures of heptameric PA channels with partially unfolded LF and EF at 4.6 and 3.1-Å resolution, respectively. The first α helix and β strand of LF and EF unfold and dock into a deep amphipathic cleft, called the α clamp, which resides at the interface of two PA monomers. The α-clamp-helix interactions exhibit structural plasticity when comparing the structures of lethal and edema toxins. EF undergoes a largescale conformational rearrangement when forming the complex with the channel. A critical loop in the PA binding interface is displaced for about 4 Å, leading to the weakening of the binding interface prior to translocation. These structures provide key insights into the molecular mechanisms of translocation-coupled protein unfolding and translocation.
Organizational Affiliation:
Department of Microbial Pathogenesis, University of Maryland, Baltimore, Baltimore, MD, 21201, USA.