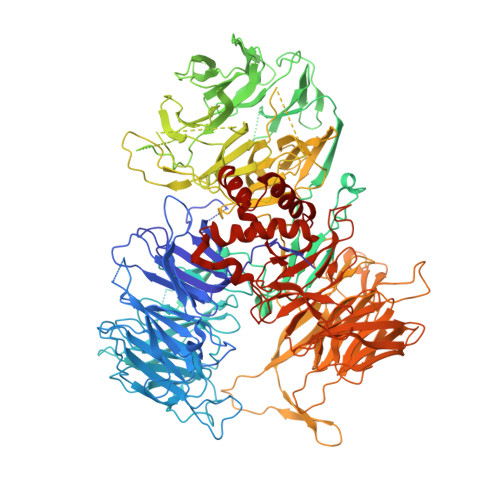
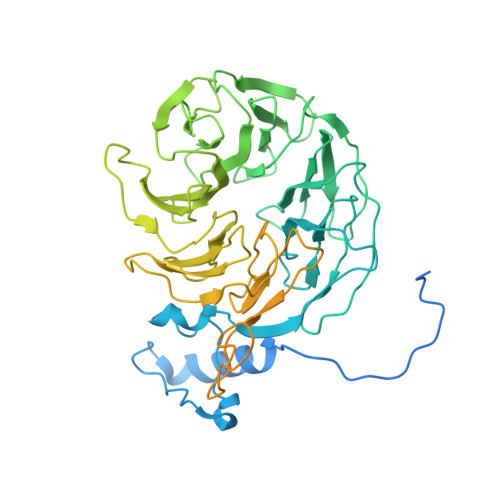
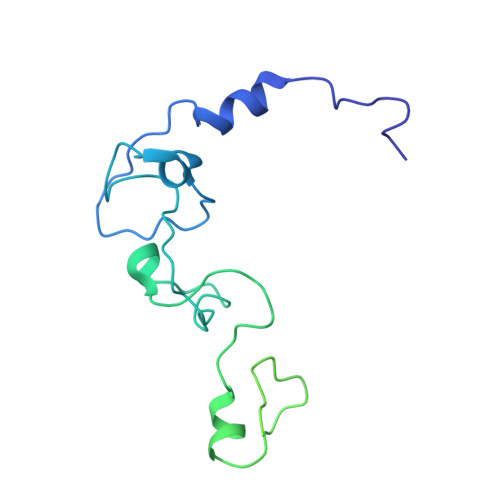
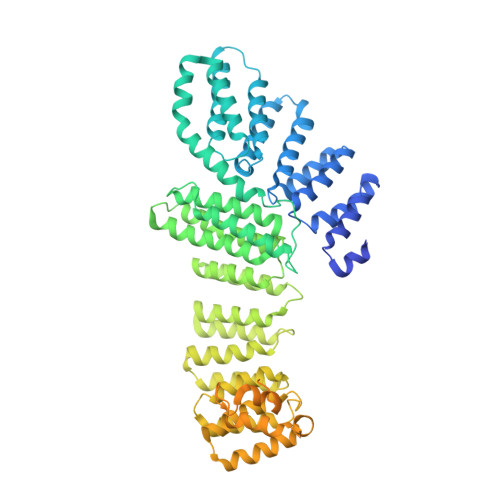
Structural Insights into the Human Pre-mRNA 3'-End Processing Machinery.
Zhang, Y., Sun, Y., Shi, Y., Walz, T., Tong, L.(2020) Mol Cell 77: 800
- PubMed: 31810758
- DOI: https://doi.org/10.1016/j.molcel.2019.11.005
- Primary Citation of Related Structures:
6URG, 6URO - PubMed Abstract:
The mammalian pre-mRNA 3'-end-processing machinery consists of cleavage and polyadenylation specificity factor (CPSF), cleavage stimulation factor (CstF), and other proteins, but the overall architecture of this machinery remains unclear. CPSF contains two functionally distinct modules: a cleavage factor (mCF) and a polyadenylation specificity factor (mPSF). Here, we have produced recombinant human CPSF and CstF and examined these factors by electron microscopy (EM). We find that mPSF is the organizational core of the machinery, while the conformations of mCF and CstF and the position of mCF relative to mPSF are highly variable. We have identified by cryo-EM a segment in CPSF100 that tethers mCF to mPSF, and we have named it the PSF interaction motif (PIM). Mutations in the PIM can abolish CPSF formation, indicating that it is a crucial contact in CPSF. We have also obtained reconstructions of mCF and CstF77 by cryo-EM, assembled around the mPSF core.
Organizational Affiliation:
Laboratory of Molecular Electron Microscopy, Rockefeller University, New York, NY 10065, USA.