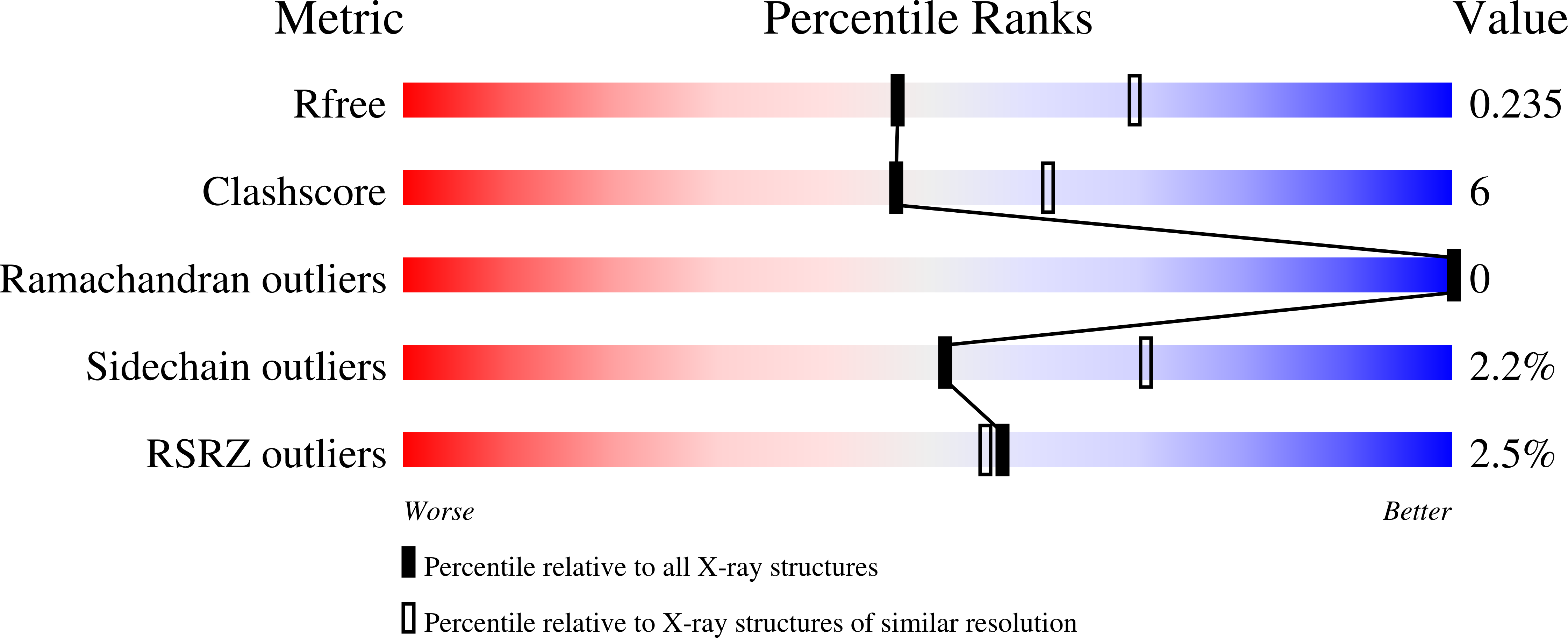
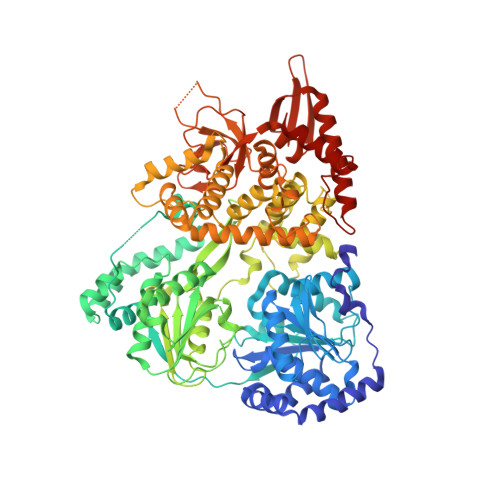
Function of Auxiliary Domains of the DEAH/RHA Helicase DHX36 in RNA Remodeling.
Srinivasan, S., Liu, Z., Chuenchor, W., Xiao, T.S., Jankowsky, E.(2020) J Mol Biol 432: 2217-2231
- PubMed: 32087197
- DOI: https://doi.org/10.1016/j.jmb.2020.02.005
- Primary Citation of Related Structures:
6UP4 - PubMed Abstract:
The DEAH/RHA helicase DHX36 has been linked to cellular RNA and DNA quadruplex structures and to AU-rich RNA elements. In vitro, DHX36 remodels DNA and RNA quadruplex structures and unwinds DNA duplexes in an ATP-dependent manner. DHX36 contains the superfamily 2 helicase core and several auxiliary domains that are conserved in orthologs of the enzyme. The role of these auxiliary domains for the enzymatic function of DHX36 is not well understood. Here, we combine structural and biochemical studies to define the function of three auxiliary domains that contact nucleic acid. We first report the crystal structure of mouse DHX36 bound to ADP. The structure reveals an overall architecture of mouse DHX36 that is similar to previously reported architectures of fly and bovine DHX36. In addition, our structure shows conformational changes that accompany stages of the ATP-binding and hydrolysis cycle. We then examine the roles of the DHX36-specific motif (DSM), the OB-fold, and a conserved β-hairpin (β-HP) in mouse DHX36 in the remodeling of RNA structures. We demonstrate and characterize RNA duplex unwinding for DHX36 and examine the remodeling of inter- and intramolecular RNA quadruplex structures. We find that the DSM not only functions as a quadruplex binding adaptor but also promotes the remodeling of RNA duplex and quadruplex structures. The OB-fold and the β-HP contribute to RNA binding. Both domains are also essential for remodeling RNA quadruplex and duplex structures. Our data reveal roles of auxiliary domains for multiple steps of the nucleic acid remodeling reactions.
Organizational Affiliation:
Center for RNA Science and Therapeutics, USA.