Impact of DNA sequences on DNA 'opening' by the Rad4/XPC nucleotide excision repair complex.
Paul, D., Mu, H., Tavakoli, A., Dai, Q., Chakraborty, S., He, C., Ansari, A., Broyde, S., Min, J.H.(2021) DNA Repair (Amst) 107: 103194-103194
- PubMed: 34428697
- DOI: https://doi.org/10.1016/j.dnarep.2021.103194
- Primary Citation of Related Structures:
6UG1 - PubMed Abstract:
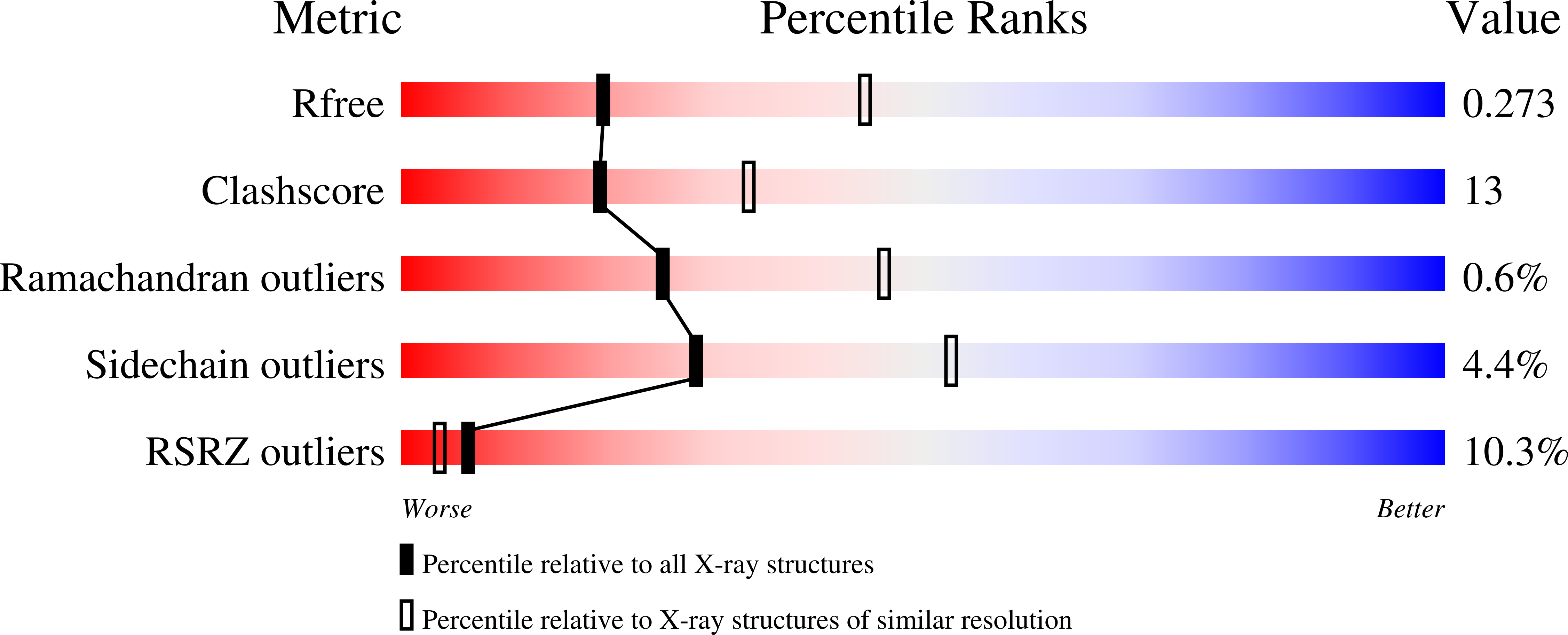
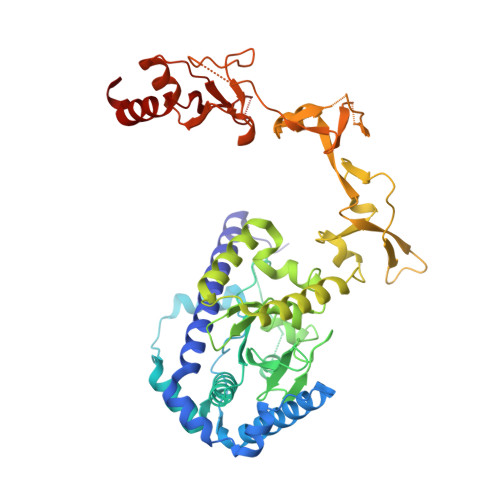
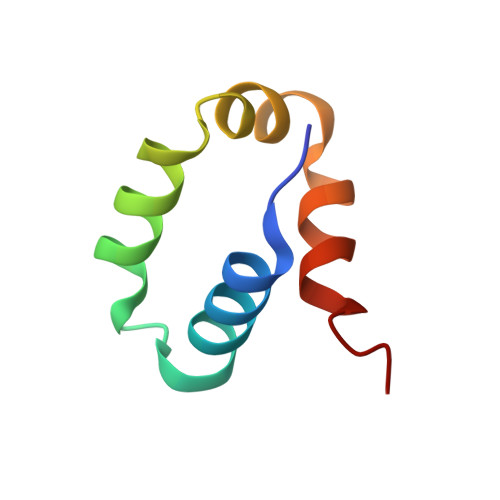
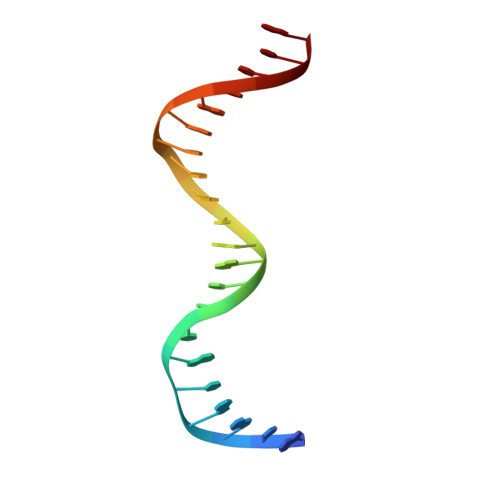
Rad4/XPC recognizes diverse DNA lesions to initiate nucleotide excision repair (NER). However, NER propensities among lesions vary widely and repair-resistant lesions are persistent and thus highly mutagenic. Rad4 recognizes repair-proficient lesions by unwinding ('opening') the damaged DNA site. Such 'opening' is also observed on a normal DNA sequence containing consecutive C/G's (CCC/GGG) when tethered to Rad4 to prevent protein diffusion. However, it was unknown if such tethering-facilitated DNA 'opening' could occur on any DNA or if certain structures/sequences would resist being 'opened'. Here, we report that DNA containing alternating C/G's (CGC/GCG) failed to be opened even when tethered; instead, Rad4 bound in a 180°-reversed manner, capping the DNA end. Fluorescence lifetime studies of DNA conformations in solution showed that CCC/GGG exhibits local pre-melting that is absent in CGC/GCG. In MD simulations, CGC/GCG failed to engage Rad4 to promote 'opening' contrary to CCC/GGG. Altogether, our study illustrates how local sequences can impact DNA recognition by Rad4/XPC and how certain DNA sites resist being 'opened' even with Rad4 held at that site indefinitely. The contrast between CCC/GGG and CGC/GCG sequences in Rad4-DNA recognition may help decipher a lesion's mutagenicity in various genomic sequence contexts to explain lesion-determined mutational hot and cold spots.
Organizational Affiliation:
Department of Chemistry and Biochemistry, Baylor University, Waco, TX, 76798, USA.