Flavonoid-based inhibitors of the Phi-class glutathione transferase from black-grass to combat multiple herbicide resistance.
Schwarz, M., Eno, R.F.M., Freitag-Pohl, S., Coxon, C.R., Straker, H.E., Wortley, D.J., Hughes, D.J., Mitchell, G., Moore, J., Cummins, I., Onkokesung, N., Brazier-Hicks, M., Edwards, R., Pohl, E., Steel, P.G.(2021) Org Biomol Chem 19: 9211-9222
- PubMed: 34643629
- DOI: https://doi.org/10.1039/d1ob01802g
- Primary Citation of Related Structures:
6TO3, 7OBO, 7ODM - PubMed Abstract:
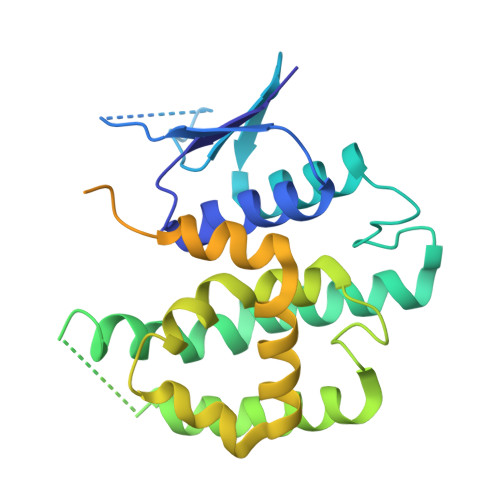
The evolution and growth of multiple-herbicide resistance (MHR) in grass weeds continues to threaten global cereal production. While various processes can contribute to resistance, earlier work has identified the phi class glutathione- S -transferase ( Am GSTF1) as a functional biomarker of MHR in black-grass ( Alopecurus myosuroides ). This study provides further insights into the role of Am GSTF1 in MHR using a combination of chemical and structural biology. Crystal structures of wild-type Am GSTF1, together with two specifically designed variants that allowed the co-crystal structure determination with glutathione and a glutathione adduct of the Am GSTF1 inhibitor 4-chloro-7-nitro-benzofurazan (NBD-Cl) were obtained. These studies demonstrated that the inhibitory activity of NBD-Cl was associated with the occlusion of the active site and the impediment of substrate binding. A search for other selective inhibitors of Am GSTF1, using ligand-fishing experiments, identified a number of flavonoids as potential ligands. Subsequent experiments using black-grass extracts discovered a specific flavonoid as a natural ligand of the recombinant enzyme. A series of related synthetic flavonoids was prepared and their binding to Am GSTF1 was investigated showing a high affinity for derivatives bearing a O -5-decyl-α-carboxylate. Molecular modelling based on high-resolution crystal structures allowed a binding pose to be defined which explained flavonoid binding specificity. Crucially, high binding affinity was linked to a reversal of the herbicide resistance phenotype in MHR black-grass. Collectively, these results present a nature-inspired new lead for the development of herbicide synergists to counteract MHR in weeds.
Organizational Affiliation:
Department of Chemistry, University of Durham, Science Laboratories, South Road, Durham, DH1 3LE, UK. p.g.steel@durham.ac.uk.