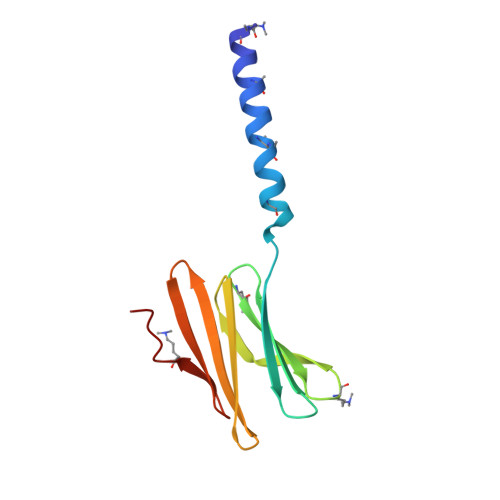
Segregated Protein-Cucurbit[7]uril Crystalline Architectures via Modulatory Peptide Tectons.
Ramberg, K.O., Guagnini, F., Engilberge, S., Wronska, M.A., Rennie, M.L., Perez, J., Crowley, P.B.(2021) Chemistry
- PubMed: 34432924
- DOI: https://doi.org/10.1002/chem.202103025
- Primary Citation of Related Structures:
6S99, 7P2H, 7P2I, 7P2J - PubMed Abstract:
One approach to protein assembly involves water-soluble supramolecular receptors that act like glues. Bionanoarchitectures directed by these scaffolds are often system-specific, with few studies investigating their customization. Herein, the modulation of cucurbituril-mediated protein assemblies through the inclusion of peptide tectons is described. Three peptides of varying length and structural order were N-terminally appended to RSL, a β-propeller building block. Each fusion protein was incorporated into crystalline architectures mediated by cucurbit[7]uril (Q7). A trimeric coiled-coil served as a spacer within a Q7-directed sheet assembly of RSL, giving rise to a layered material of varying porosity. Within the spacer layers, the coiled-coils were dynamic. This result prompted consideration of intrinsically disordered peptides (IDPs) as modulatory tectons. Similar to the coiled-coil, a mussel adhesion peptide (Mefp) also acted as a spacer between protein-Q7 sheets. In contrast, the fusion of a nucleoporin peptide (Nup) to RSL did not recapitulate the sheet assembly. Instead, a Q7-directed cage was adopted, within which disordered Nup peptides were partially "captured" by Q7 receptors. IDP capture occurred by macrocycle recognition of an intrapeptide Phe-Gly motif in which the benzyl group was encapsulated by Q7. The modularity of these protein-cucurbituril architectures adds a new dimension to macrocycle-mediated protein assembly. Segregated protein crystals, with alternating layers of high and low porosity, could provide a basis for new types of materials.
Organizational Affiliation:
School of Chemistry, National University of Ireland Galway, University Road, Galway, H91 TK33, Ireland.